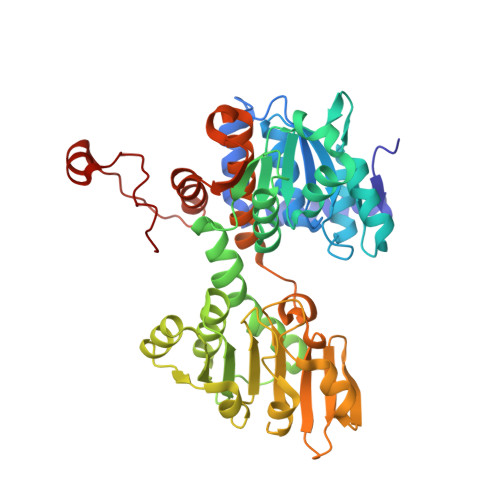
Discovery and structural analyses of S-adenosyl-L-homocysteine hydrolase inhibitors based on non-adenosine analogs.
Nakao, A., Suzuki, H., Ueno, H., Iwasaki, H., Setsuta, T., Kashima, A., Sunada, S.(2015) Bioorg Med Chem 23: 4952-4969
- PubMed: 26037610 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2015.05.018
- Primary Citation Related Structures:
4YVF - PubMed Abstract:
Optimization of a new series of S-adenosyl-L-homocysteine hydrolase (AdoHcyase) inhibitors based on non-adenosine analogs led to very potent compounds 14n, 18a, and 18b with IC50 values of 13 ± 3, 5.0 ± 2.0, and 8.5 ± 3.1 nM, respectively. An X-ray crystal structure of AdoHcyase with NAD(+) and 18a showed a novel open form co-crystal structure. 18a in the co-crystals formed intramolecular eight membered ring hydrogen bond formations. A single crystal X-ray structure of 14n also showed an intramolecular eight-membered ring hydrogen bond interaction.
- Research Division, Mitsubishi Tanabe Pharma Corporation, 1000, Kamoshida-cho, Aoba-ku, Yokohama 227-0033, Japan. Electronic address: Nakao.Akira@mc.mt-pharma.co.jp.
Organizational Affiliation: