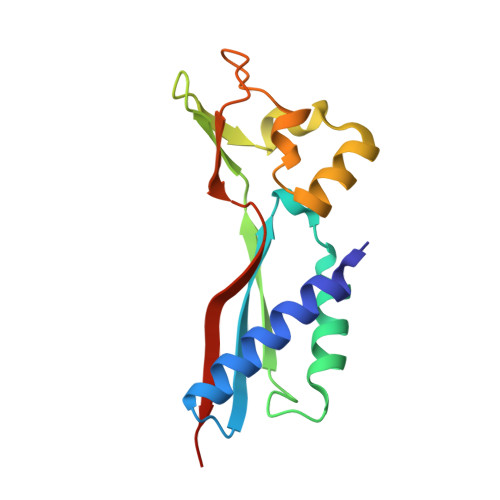
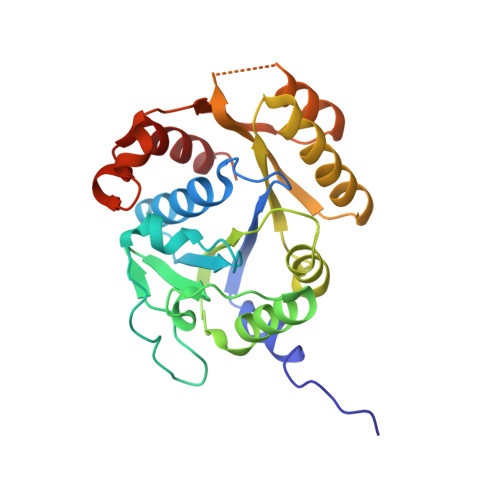
Structural basis of a Ni acquisition cycle for [NiFe] hydrogenase by Ni-metallochaperone HypA and its enhancer
Watanabe, S., Kawashima, T., Nishitani, Y., Kanai, T., Wada, T., Inaba, K., Atomi, H., Imanaka, T., Miki, K.(2015) Proc Natl Acad Sci U S A 112: 7701-7706
- PubMed: 26056269 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1503102112
- Primary Citation Related Structures:
5AUN, 5AUO, 5AUP, 5AUQ - PubMed Abstract:
The Ni atom at the catalytic center of [NiFe] hydrogenases is incorporated by a Ni-metallochaperone, HypA, and a GTPase/ATPase, HypB. We report the crystal structures of the transient complex formed between HypA and ATPase-type HypB (HypBAT) with Ni ions. Transient association between HypA and HypBAT is controlled by the ATP hydrolysis cycle of HypBAT, which is accelerated by HypA. Only the ATP-bound form of HypBAT can interact with HypA and induces drastic conformational changes of HypA. Consequently, upon complex formation, a conserved His residue of HypA comes close to the N-terminal conserved motif of HypA and forms a Ni-binding site, to which a Ni ion is bound with a nearly square-planar geometry. The Ni binding site in the HypABAT complex has a nanomolar affinity (Kd = 7 nM), which is in contrast to the micromolar affinity (Kd = 4 µM) observed with the isolated HypA. The ATP hydrolysis and Ni binding cause conformational changes of HypBAT, affecting its association with HypA. These findings indicate that HypA and HypBAT constitute an ATP-dependent Ni acquisition cycle for [NiFe]-hydrogenase maturation, wherein HypBAT functions as a metallochaperone enhancer and considerably increases the Ni-binding affinity of HypA.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan; Institute of Multidisciplinary Research for Advanced Materials, Tohoku University, Katahira 2-1-1, Aoba-ku, Sendai 980-8577, Japan;
Organizational Affiliation: