Studies on Aryl-Substituted Phenylalanines: Synthesis, Activity, and Different Binding Modes at AMPA Receptors.
Szymanska, E., Frydenvang, K., Pickering, D.S., Krintel, C., Nielsen, B., Kooshki, A., Zachariassen, L.G., Olsen, L., Kastrup, J.S., Johansen, T.N.(2016) J Med Chem 59: 448-461
- PubMed: 26653877 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01666
- Primary Citation Related Structures:
5CBR, 5CBS - PubMed Abstract:
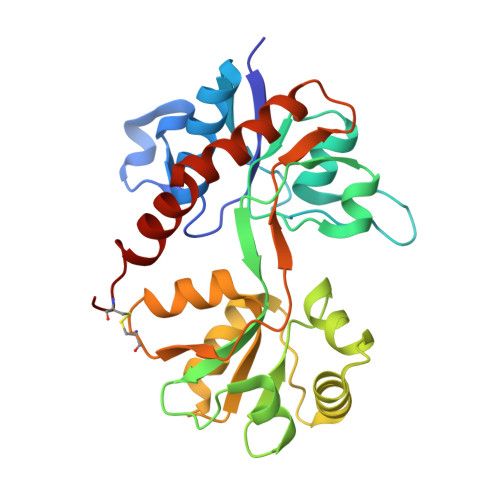
A series of racemic aryl-substituted phenylalanines was synthesized and evaluated in vitro at recombinant rat GluA1-3, at GluK1-3, and at native AMPA receptors. The individual enantiomers of two target compounds, (RS)-2-amino-3-(3,4-dichloro-5-(5-hydroxypyridin-3-yl)phenyl)propanoic acid 37 and (RS)-2-amino-3-(3'-hydroxybiphenyl-3-yl)propanoic acid 38, were characterized. (S)-37 and (R)-38 were identified as the only biologically active isomers, both being antagonists at GluA2 receptors with Kb of 1.80 and 3.90 μM, respectively. To address this difference in enantiopharmacology, not previously seen for amino acid-based AMPA receptor antagonists, X-ray crystal structures of both eutomers in complex with the GluA2 ligand binding domain were solved. The cocrystal structures of (S)-37 and (R)-38 showed similar interactions of the amino acid parts but unexpected and different orientations and interactions of the biaromatic parts of the ligands inside the binding site, with (R)-38 having a binding mode not previously identified for amino acid-based antagonists.
- Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen , 2100 Copenhagen, Denmark.
Organizational Affiliation: