Development of an in-vivo active reversible butyrylcholinesterase inhibitor.
Kosak, U., Brus, B., Knez, D., Sink, R., Zakelj, S., Trontelj, J., Pislar, A., Slenc, J., Gobec, M., Zivin, M., Tratnjek, L., Perse, M., Saat, K., Podkowa, A., Filipek, B., Nachon, F., Brazzolotto, X., Wieckowska, A., Malawska, B., Stojan, J., Rascan, I.M., Kos, J., Coquelle, N., Colletier, J.P., Gobec, S.(2016) Sci Rep 6: 39495-39495
- PubMed: 28000737 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep39495
- Primary Citation Related Structures:
5DYT, 5DYW, 5DYY - PubMed Abstract:
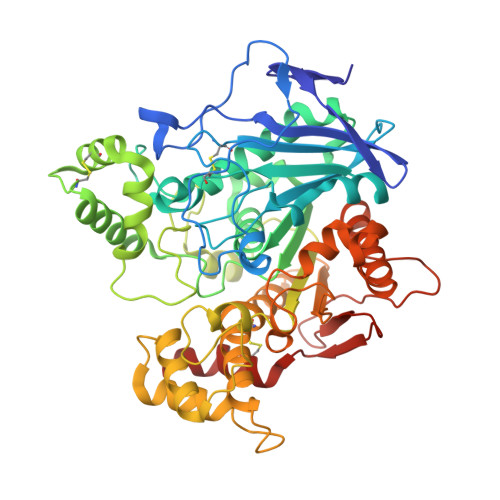
Alzheimer's disease (AD) is characterized by severe basal forebrain cholinergic deficit, which results in progressive and chronic deterioration of memory and cognitive functions. Similar to acetylcholinesterase, butyrylcholinesterase (BChE) contributes to the termination of cholinergic neurotransmission. Its enzymatic activity increases with the disease progression, thus classifying BChE as a viable therapeutic target in advanced AD. Potent, selective and reversible human BChE inhibitors were developed. The solved crystal structure of human BChE in complex with the most potent inhibitor reveals its binding mode and provides the molecular basis of its low nanomolar potency. Additionally, this compound is noncytotoxic and has neuroprotective properties. Furthermore, this inhibitor moderately crosses the blood-brain barrier and improves memory, cognitive functions and learning abilities of mice in a model of the cholinergic deficit that characterizes AD, without producing acute cholinergic adverse effects. Our study provides an advanced lead compound for developing drugs for alleviating symptoms caused by cholinergic hypofunction in advanced AD.
- Faculty of Pharmacy, University of Ljubljana, Aškerčeva 7, 1000 Ljubljana, Slovenia.
Organizational Affiliation: