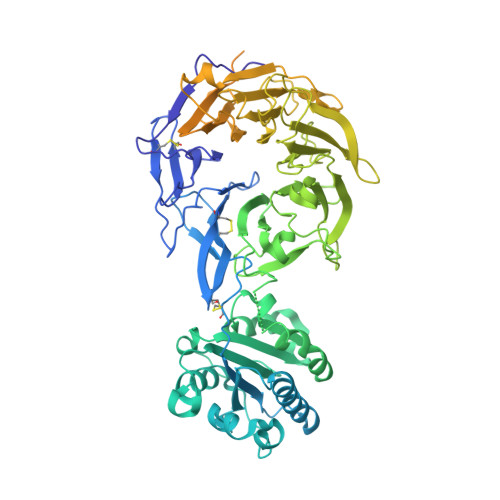
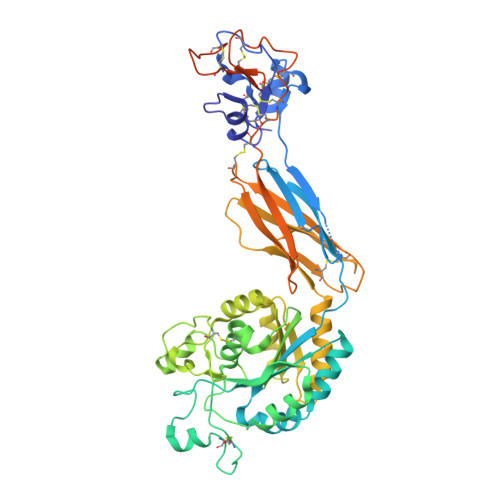
Leukocyte integrin alpha L beta 2 headpiece structures: The alpha I domain, the pocket for the internal ligand, and concerted movements of its loops.
Sen, M., Springer, T.A.(2016) Proc Natl Acad Sci U S A 113: 2940-2945
- PubMed: 26936951 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1601379113
- Primary Citation Related Structures:
5E6R, 5E6S, 5E6U, 5E6V, 5E6W, 5E6X, 5ES4 - PubMed Abstract:
High-resolution crystal structures of the headpiece of lymphocyte function-associated antigen-1 (integrin αLβ2) reveal how the αI domain interacts with its platform formed by the α-subunit β-propeller and β-subunit βI domains. The αLβ2 structures compared with αXβ2 structures show that the αI domain, tethered through its N-linker and a disulfide to a stable β-ribbon pillar near the center of the platform, can undergo remarkable pivoting and tilting motions that appear buffered by N-glycan decorations that differ between αL and αX subunits. Rerefined β2 integrin structures reveal details including pyroglutamic acid at the β2 N terminus and bending within the EGF1 domain. Allostery is relayed to the αI domain by an internal ligand that binds to a pocket at the interface between the β-propeller and βI domains. Marked differences between the αL and αX subunit β-propeller domains concentrate near the binding pocket and αI domain interfaces. Remarkably, movement in allostery in the βI domain of specificity determining loop 1 (SDL1) causes concerted movement of SDL2 and thereby tightens the binding pocket for the internal ligand.
- Program in Cellular and Molecular Medicine, Children's Hospital Boston, and Departments of Biological Chemistry and Molecular Pharmacology and of Medicine, Harvard Medical School, Boston, MA 02115; Department of Biology and Biochemistry, University of Houston, Houston, TX 77204.
Organizational Affiliation: