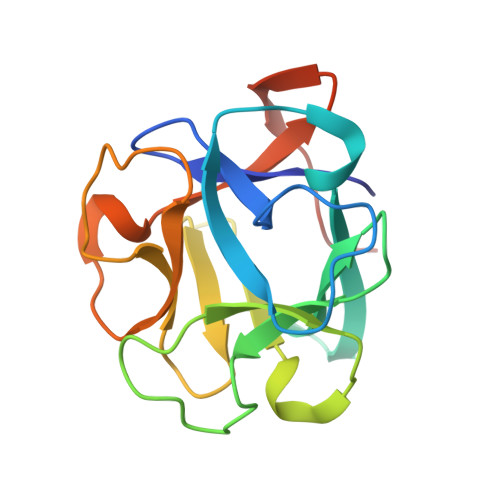
A Multivalent Marine Lectin from Crenomytilus grayanus Possesses Anti-cancer Activity through Recognizing Globotriose Gb3
Liao, J.-H., Chien, C.-T., Wu, H.-Y., Huang, K.-F., Wang, I., Ho, M.-R., Tu, I.-F., Lee, I.-M., Li, W., Shih, Y.-L., Wu, C.-Y., Lukyanov, P.A., Hsu, S.D., Wu, S.-H.(2016) J Am Chem Soc 138: 4787-4795
- PubMed: 27010847
- DOI: https://doi.org/10.1021/jacs.6b00111
- Primary Citation of Related Structures:
5F8S, 5F8W, 5F8Y, 5F90 - PubMed Abstract:
In this study, we report the structure and function of a lectin from the sea mollusk Crenomytilus grayanus collected from the sublittoral zone of Peter the Great Bay of the Sea of Japan. The crystal structure of C. grayanus lectin (CGL) was solved to a resolution of 1.08 Å, revealing a β-trefoil fold that dimerizes into a dumbbell-shaped quaternary structure. Analysis of the crystal CGL structures bound to galactose, galactosamine, and globotriose Gb3 indicated that each CGL can bind three ligands through a carbohydrate-binding motif involving an extensive histidine- and water-mediated hydrogen bond network. CGL binding to Gb3 is further enhanced by additional side-chain-mediated hydrogen bonds in each of the three ligand-binding sites. NMR titrations revealed that the three binding sites have distinct microscopic affinities toward galactose and galactosamine. Cell viability assays showed that CGL recognizes Gb3 on the surface of breast cancer cells, leading to cell death. Our findings suggest the use of this lectin in cancer diagnosis and treatment.
Organizational Affiliation:
Institute of Biological Chemistry, Academia Sinica , Taipei 11529, Taiwan.