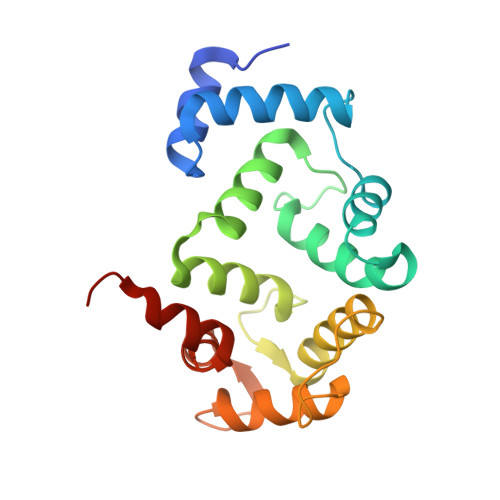
Biophysical and functional characterization of hippocalcin mutants responsible for human dystonia.
Helassa, N., Antonyuk, S.V., Lian, L.Y., Haynes, L.P., Burgoyne, R.D.(2017) Hum Mol Genet 26: 2426-2435
- PubMed: 28398555 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/hmg/ddx133
- Primary Citation Related Structures:
5G4P, 5G58, 5M6C - PubMed Abstract:
Dystonia is a neurological movement disorder that forces the body into twisting, repetitive movements or sometimes painful abnormal postures. With the advent of next-generation sequencing technologies, the homozygous mutations T71N and A190T in the neuronal calcium sensor (NCS) hippocalcin were identified as the genetic cause of primary isolated dystonia (DYT2 dystonia). However, the effect of these mutations on the physiological role of hippocalcin has not yet been elucidated. Using a multidisciplinary approach, we demonstrated that hippocalcin oligomerises in a calcium-dependent manner and binds to voltage-gated calcium channels. Mutations T71N and A190T in hippocalcin did not affect stability, calcium-binding affinity or translocation to cellular membranes (Ca2+/myristoyl switch). We obtained the first crystal structure of hippocalcin and alignment with other NCS proteins showed significant variability in the orientation of the C-terminal part of the molecule, the region expected to be important for target binding. We demonstrated that the disease-causing mutations did not affect the structure of the protein, however both mutants showed a defect in oligomerisation. In addition, we observed an increased calcium influx in KCl-depolarised cells expressing mutated hippocalcin, mostly driven by N-type voltage-gated calcium channels. Our data demonstrate that the dystonia-causing mutations strongly affect hippocalcin cellular functions which suggest a central role for perturbed calcium signalling in DYT2 dystonia.
- Department of Cellular and Molecular Physiology, Institute of Translational Medicine, University of Liverpool, Liverpool L69?3BX, UK.
Organizational Affiliation: