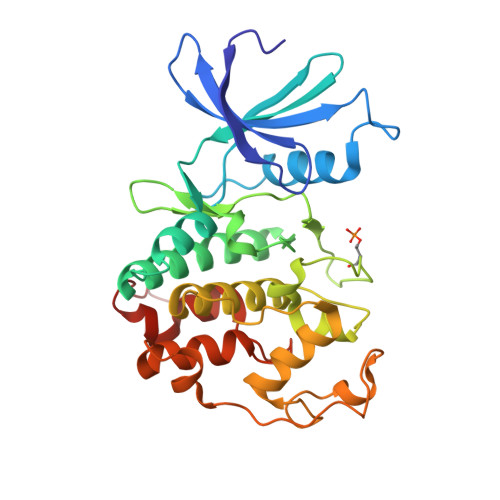
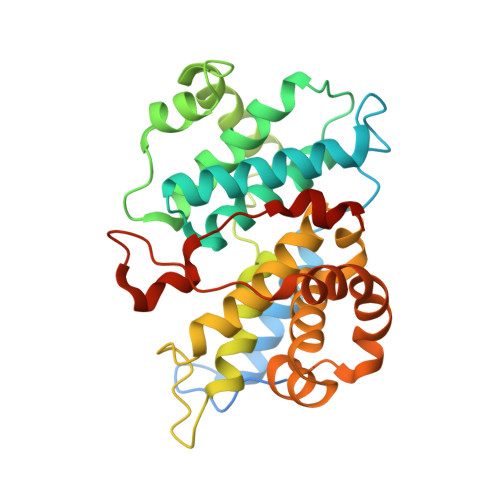
Spectrum and Degree of CDK Drug Interactions Predicts Clinical Performance.
Chen, P., Lee, N.V., Hu, W., Xu, M., Ferre, R.A., Lam, H., Bergqvist, S., Solowiej, J., Diehl, W., He, Y.A., Yu, X., Nagata, A., VanArsdale, T., Murray, B.W.(2016) Mol Cancer Ther 15: 2273-2281
- PubMed: 27496135 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-16-0300
- Primary Citation Related Structures:
5L2I, 5L2S, 5L2T, 5L2W - PubMed Abstract:
Therapeutically targeting aberrant intracellular kinase signaling is attractive from a biological perspective but drug development is often hindered by toxicities and inadequate efficacy. Predicting drug behaviors using cellular and animal models is confounded by redundant kinase activities, a lack of unique substrates, and cell-specific signaling networks. Cyclin-dependent kinase (CDK) drugs exemplify this phenomenon because they are reported to target common processes yet have distinct clinical activities. Tumor cell studies of ATP-competitive CDK drugs (dinaciclib, AG-024322, abemaciclib, palbociclib, ribociclib) indicate similar pharmacology while analyses in untransformed cells illuminates significant differences. To resolve this apparent disconnect, drug behaviors are described at the molecular level. Nonkinase binding studies and kinome interaction analysis (recombinant and endogenous kinases) reveal that proteins outside of the CDK family appear to have little role in dinaciclib/palbociclib/ribociclib pharmacology, may contribute for abemaciclib, and confounds AG-024322 analysis. CDK2 and CDK6 cocrystal structures with the drugs identify the molecular interactions responsible for potency and kinase selectivity. Efficient drug binding to the unique hinge architecture of CDKs enables selectivity toward most of the human kinome. Selectivity between CDK family members is achieved through interactions with nonconserved elements of the ATP-binding pocket. Integrating clinical drug exposures into the analysis predicts that both palbociclib and ribociclib are CDK4/6 inhibitors, abemaciclib inhibits CDK4/6/9, and dinaciclib is a broad-spectrum CDK inhibitor (CDK2/3/4/6/9). Understanding the molecular components of potency and selectivity also facilitates rational design of future generations of kinase-directed drugs. Mol Cancer Ther; 15(10); 2273-81. ©2016 AACR.
- Oncology Medicinal Chemistry, Pfizer Worldwide Research and Development, San Diego, California.
Organizational Affiliation: