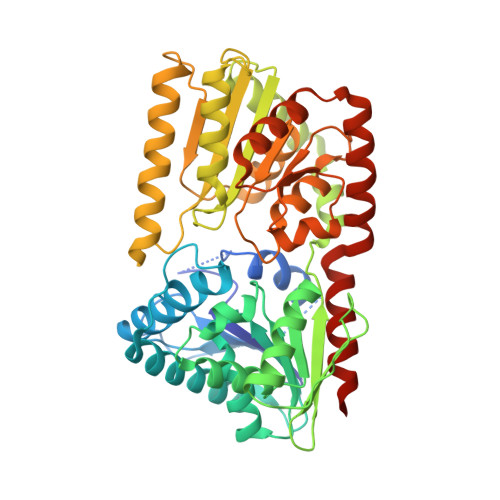
Mycobacterial OtsA Structures Unveil Substrate Preference Mechanism and Allosteric Regulation by 2-Oxoglutarate and 2-Phosphoglycerate.
Mendes, V., Acebron-Garcia-de-Eulate, M., Verma, N., Blaszczyk, M., Dias, M.V.B., Blundell, T.L.(2019) mBio 10
- PubMed: 31772052 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.02272-19
- Primary Citation Related Structures:
5JIJ, 5JIO, 5K41, 5K42, 5K44, 5K5C, 5L3K - PubMed Abstract:
Trehalose is an essential disaccharide for mycobacteria and a key constituent of several cell wall glycolipids with fundamental roles in pathogenesis. Mycobacteria possess two pathways for trehalose biosynthesis. However, only the OtsAB pathway was found to be essential in Mycobacterium tuberculosis , with marked growth and virulence defects of OtsA mutants and strict essentiality of OtsB2. Here, we report the first mycobacterial OtsA structures from Mycobacterium thermoresistibile in both apo and ligand-bound forms. Structural information reveals three key residues in the mechanism of substrate preference that were further confirmed by site-directed mutagenesis. Additionally, we identify 2-oxoglutarate and 2-phosphoglycerate as allosteric regulators of OtsA. The structural analysis in this work strongly contributed to define the mechanisms for feedback inhibition, show different conformational states of the enzyme, and map a new allosteric site. IMPORTANCE Mycobacterial infections are a significant source of mortality worldwide, causing millions of deaths annually. Trehalose is a multipurpose disaccharide that plays a fundamental structural role in these organisms as a component of mycolic acids, a molecular hallmark of the cell envelope of mycobacteria. Here, we describe the first mycobacterial OtsA structures. We show mechanisms of substrate preference and show that OtsA is regulated allosterically by 2-oxoglutarate and 2-phosphoglycerate at an interfacial site. These results identify a new allosteric site and provide insight on the regulation of trehalose synthesis through the OtsAB pathway in mycobacteria.
- Department of Biochemistry, University of Cambridge, Cambridge, United Kingdom vgm23@cam.ac.uk tom@cryst.bioc.cam.ac.uk.
Organizational Affiliation: