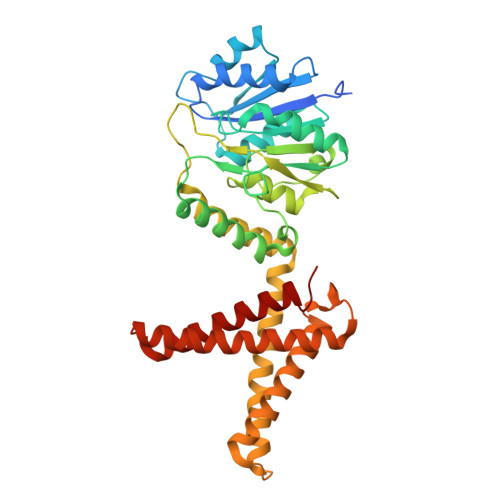
Structural basis for dolichylphosphate mannose biosynthesis.
Gandini, R., Reichenbach, T., Tan, T.C., Divne, C.(2017) Nat Commun 8: 120-120
- PubMed: 28743912 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-00187-2
- Primary Citation Related Structures:
5MLZ, 5MM0, 5MM1 - PubMed Abstract:
Protein glycosylation is a critical protein modification. In biogenic membranes of eukaryotes and archaea, these reactions require activated mannose in the form of the lipid conjugate dolichylphosphate mannose (Dol-P-Man). The membrane protein dolichylphosphate mannose synthase (DPMS) catalyzes the reaction whereby mannose is transferred from GDP-mannose to the dolichol carrier Dol-P, to yield Dol-P-Man. Failure to produce or utilize Dol-P-Man compromises organism viability, and in humans, several mutations in the human dpm1 gene lead to congenital disorders of glycosylation (CDG). Here, we report three high-resolution crystal structures of archaeal DPMS from Pyrococcus furiosus, in complex with nucleotide, donor, and glycolipid product. The structures offer snapshots along the catalytic cycle, and reveal how lipid binding couples to movements of interface helices, metal binding, and acceptor loop dynamics to control critical events leading to Dol-P-Man synthesis. The structures also rationalize the loss of dolichylphosphate mannose synthase function in dpm1-associated CDG.The generation of glycolipid dolichylphosphate mannose (Dol-P-Man) is a critical step for protein glycosylation and GPI anchor synthesis. Here the authors report the structure of dolichylphosphate mannose synthase in complex with bound nucleotide and donor to provide insight into the mechanism of Dol-P-Man synthesis.
- School of Biotechnology, KTH Royal Institute of Technology, S-10691, Stockholm, Sweden.
Organizational Affiliation: