Crystal structure of DC8E8 Fab in the complex with a 14-mer tau peptide at pH 8.5
Skrabana, R., Novak, M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Microtubule-associated protein tau | 14 | Homo sapiens | Mutation(s): 0 | 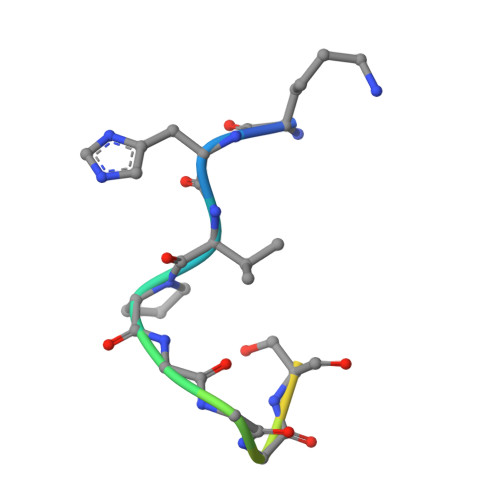 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P10636 GTEx: ENSG00000186868 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P10636 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab of monoclonal antibody, heavy chain | B [auth H] | 221 | Mus musculus | Mutation(s): 0 | 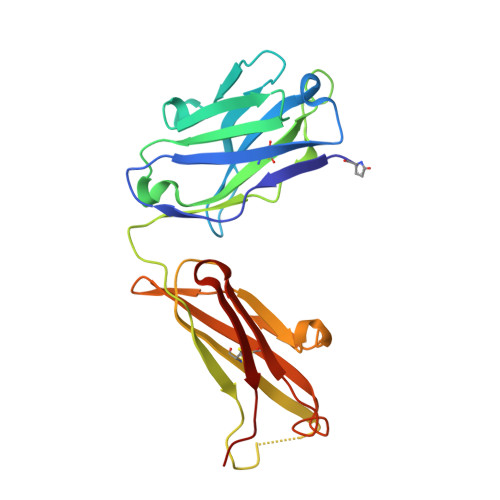 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab of monoclonal antibody, light chain | C [auth L] | 218 | Mus musculus | Mutation(s): 0 | 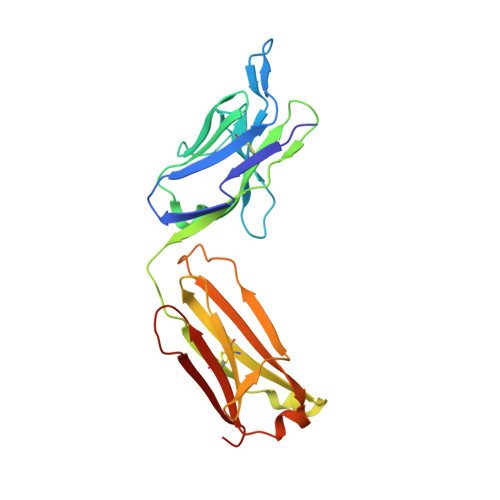 |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PGE Download:Ideal Coordinates CCD File | E [auth H], F [auth H], G [auth L], H [auth L] | TRIETHYLENE GLYCOL C6 H14 O4 ZIBGPFATKBEMQZ-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| PCA Query on PCA | B [auth H] | L-PEPTIDE LINKING | C5 H7 N O3 |  | GLN |
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900006 Query on PRD_900006 | D [auth B] | trehalose | Oligosaccharide / Nutrient |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 58.728 | α = 90 |
| b = 60.128 | β = 109.25 |
| c = 69.315 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| PHASER | phasing |