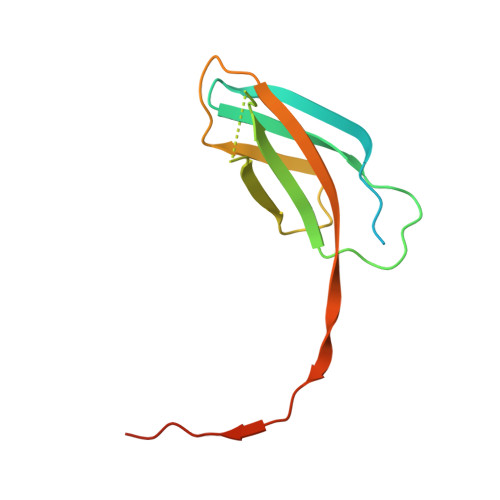
Structural basis of Tie2 activation and Tie2/Tie1 heterodimerization.
Leppanen, V.M., Saharinen, P., Alitalo, K.(2017) Proc Natl Acad Sci U S A 114: 4376-4381
- PubMed: 28396439 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1616166114
- Primary Citation Related Structures:
5MYA, 5MYB, 5N06 - PubMed Abstract:
The endothelial cell (EC)-specific receptor tyrosine kinases Tie1 and Tie2 are necessary for the remodeling and maturation of blood and lymphatic vessels. Angiopoietin-1 (Ang1) growth factor is a Tie2 agonist, whereas Ang2 functions as a context-dependent agonist/antagonist. The orphan receptor Tie1 modulates Tie2 activation, which is induced by association of angiopoietins with Tie2 in cis and across EC-EC junctions in trans Except for the binding of the C-terminal angiopoietin domains to the Tie2 ligand-binding domain, the mechanisms for Tie2 activation are poorly understood. We report here the structural basis of Ang1-induced Tie2 dimerization in cis and provide mechanistic insights on Ang2 antagonism, Tie1/Tie2 heterodimerization, and Tie2 clustering. We find that Ang1-induced Tie2 dimerization and activation occurs via the formation of an intermolecular β-sheet between the membrane-proximal (third) Fibronectin type III domains (Fn3) of Tie2. The structures of Tie2 and Tie1 Fn3 domains are similar and compatible with Tie2/Tie1 heterodimerization by the same mechanism. Mutagenesis of the key interaction residues of Tie2 and Tie1 Fn3 domains decreased Ang1-induced Tie2 phosphorylation and increased the basal phosphorylation of Tie1, respectively. Furthermore, the Tie2 structures revealed additional interactions between the Fn 2 (Fn2) domains that coincide with a mutation of Tie2 in primary congenital glaucoma that leads to defective Tie2 clustering and junctional localization. Mutagenesis of the Fn2-Fn2 interface increased the basal phosphorylation of Tie2, suggesting that the Fn2 interactions are essential in preformed Tie2 oligomerization. The interactions of the membrane-proximal domains could provide new targets for modulation of Tie receptor activity.
- Wihuri Research Institute, Biomedicum Helsinki, Haartmaninkatu 8, 00290 Helsinki, Finland; kari.alitalo@helsinki.fi veli-matti.leppanen@helsinki.fi.
Organizational Affiliation: