Thio-Linked UDP-Peptide Conjugates as O-GlcNAc Transferase Inhibitors.
Rafie, K., Gorelik, A., Trapannone, R., Borodkin, V.S., van Aalten, D.M.F.(2018) Bioconjug Chem 29: 1834-1840
- PubMed: 29723473 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.bioconjchem.8b00194
- Primary Citation Related Structures:
5NPR, 5NPS - PubMed Abstract:
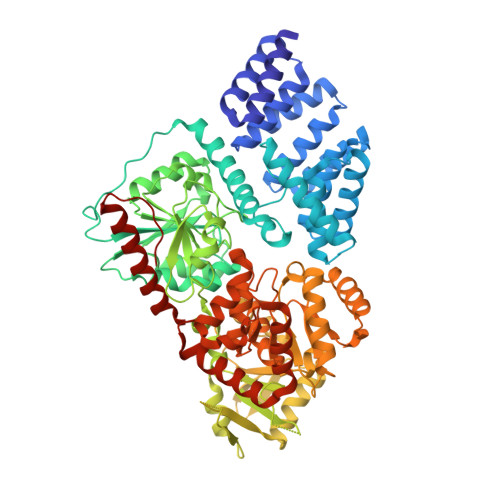
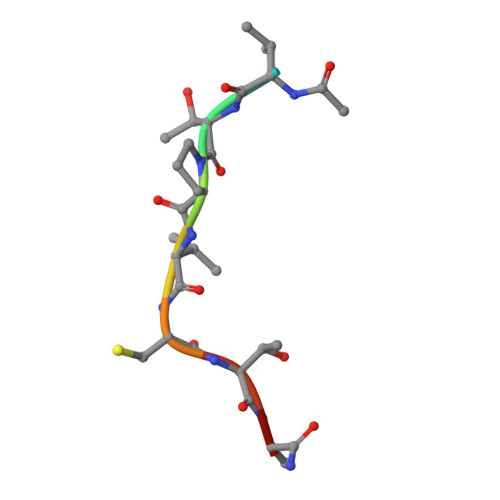
O-GlcNAc transferase (OGT) is an essential glycosyltransferase that installs the O-GlcNAc post-translational modification on the nucleocytoplasmic proteome. We report the development of S-linked UDP-peptide conjugates as potent bisubstrate OGT inhibitors. These compounds were assembled in a modular fashion by photoinitiated thiol-ene conjugation of allyl-UDP and optimal acceptor peptides in which the acceptor serine was replaced with cysteine. The conjugate VTPVC(S-propyl-UDP)TA ( K i = 1.3 μM) inhibits the OGT activity in HeLa cell lysates. Linear fusions of this conjugate with cell penetrating peptides were explored as prototypes of cell-penetrant OGT inhibitors. A crystal structure of human OGT with the inhibitor revealed mimicry of the interactions seen in the pseudo-Michaelis complex. Furthermore, a fluorophore-tagged derivative of the inhibitor works as a high affinity probe in a fluorescence polarimetry hOGT assay.
- Division of Gene Regulation and Expression, School of Life Sciences , University of Dundee , DD1 5EH Dundee , U.K.
Organizational Affiliation: