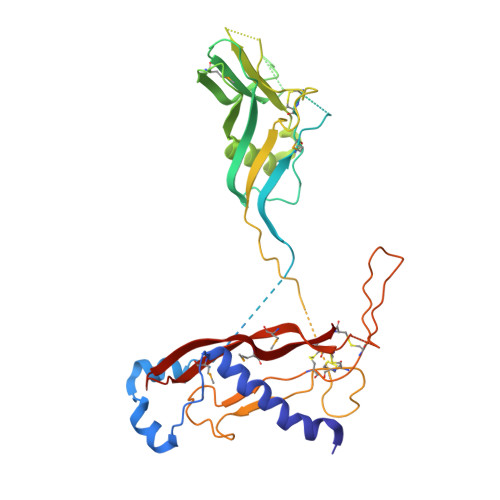
Structure of the human myostatin precursor and determinants of growth factor latency.
Cotton, T.R., Fischer, G., Wang, X., McCoy, J.C., Czepnik, M., Thompson, T.B., Hyvonen, M.(2018) EMBO J 37: 367-383
- PubMed: 29330193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.201797883
- Primary Citation Related Structures:
5NTU, 5NXS - PubMed Abstract:
Myostatin, a key regulator of muscle mass in vertebrates, is biosynthesised as a latent precursor in muscle and is activated by sequential proteolysis of the pro-domain. To investigate the molecular mechanism by which pro-myostatin remains latent, we have determined the structure of unprocessed pro-myostatin and analysed the properties of the protein in its different forms. Crystal structures and SAXS analyses show that pro-myostatin adopts an open, V-shaped structure with a domain-swapped arrangement. The pro-mature complex, after cleavage of the furin site, has significantly reduced activity compared with the mature growth factor and persists as a stable complex that is resistant to the natural antagonist follistatin. The latency appears to be conferred by a number of distinct features that collectively stabilise the interaction of the pro-domains with the mature growth factor, enabling a regulated stepwise activation process, distinct from the prototypical pro-TGF-β1. These results provide a basis for understanding the effect of missense mutations in pro-myostatin and pave the way for the design of novel myostatin inhibitors.
- Department of Biochemistry, University of Cambridge, Cambridge, UK.
Organizational Affiliation: