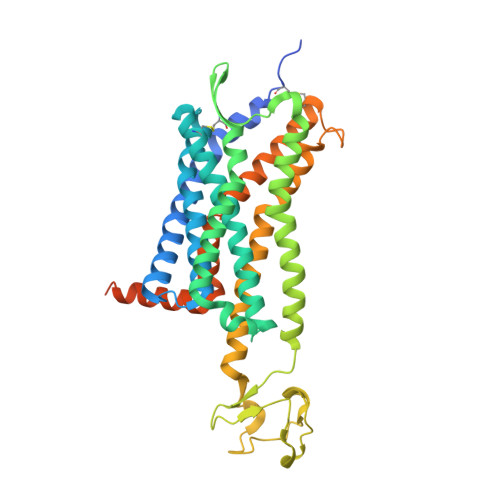
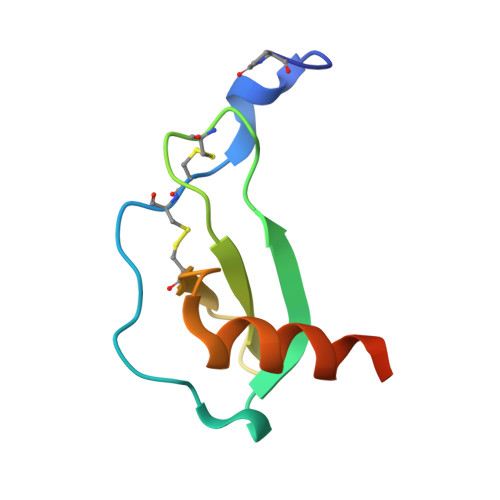
Structure of CC Chemokine Receptor 5 with a Potent Chemokine Antagonist Reveals Mechanisms of Chemokine Recognition and Molecular Mimicry by HIV.
Zheng, Y., Han, G.W., Abagyan, R., Wu, B., Stevens, R.C., Cherezov, V., Kufareva, I., Handel, T.M.(2017) Immunity 46: 1005-1017.e5
- PubMed: 28636951 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2017.05.002
- Primary Citation Related Structures:
5UIW - PubMed Abstract:
CCR5 is the primary chemokine receptor utilized by HIV to infect leukocytes, whereas CCR5 ligands inhibit infection by blocking CCR5 engagement with HIV gp120. To guide the design of improved therapeutics, we solved the structure of CCR5 in complex with chemokine antagonist [5P7]CCL5. Several structural features appeared to contribute to the anti-HIV potency of [5P7]CCL5, including the distinct chemokine orientation relative to the receptor, the near-complete occupancy of the receptor binding pocket, the dense network of intermolecular hydrogen bonds, and the similarity of binding determinants with the FDA-approved HIV inhibitor Maraviroc. Molecular modeling indicated that HIV gp120 mimicked the chemokine interaction with CCR5, providing an explanation for the ability of CCR5 to recognize diverse ligands and gp120 variants. Our findings reveal that structural plasticity facilitates receptor-chemokine specificity and enables exploitation by HIV, and provide insight into the design of small molecule and protein inhibitors for HIV and other CCR5-mediated diseases.
- University of California, San Diego, Skaggs School of Pharmacy and Pharmaceutical Sciences, La Jolla, CA 92093, USA.
Organizational Affiliation: