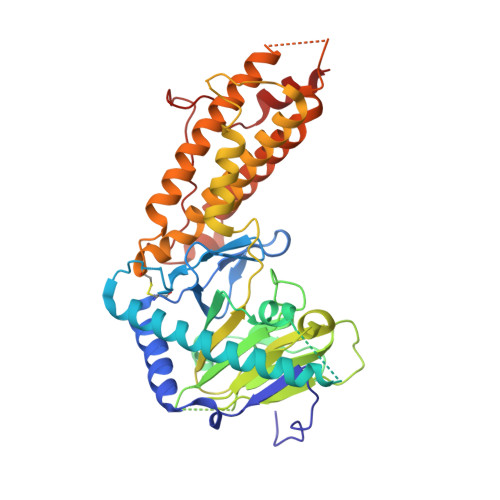
Structural insights into FTO's catalytic mechanism for the demethylation of multiple RNA substrates.
Zhang, X., Wei, L.H., Wang, Y., Xiao, Y., Liu, J., Zhang, W., Yan, N., Amu, G., Tang, X., Zhang, L., Jia, G.(2019) Proc Natl Acad Sci U S A 116: 2919-2924
- PubMed: 30718435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1820574116
- Primary Citation Related Structures:
5ZMD - PubMed Abstract:
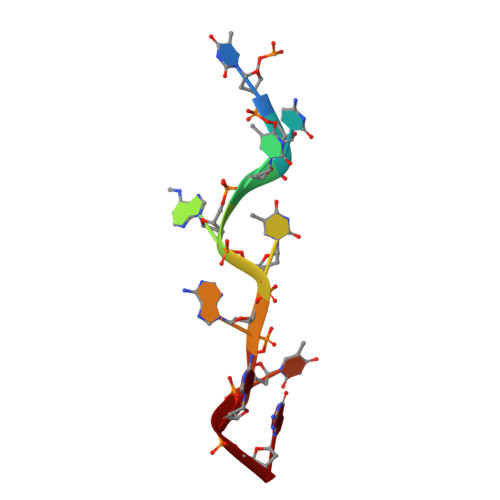
FTO demethylates internal N 6 -methyladenosine (m 6 A) and N 6 ,2'- O -dimethyladenosine (m 6 A m ; at the cap +1 position) in mRNA, m 6 A and m 6 A m in snRNA, and N 1 -methyladenosine (m 1 A) in tRNA in vivo, and in vitro evidence supports that it can also demethylate N 6 -methyldeoxyadenosine (6mA), 3-methylthymine (3mT), and 3-methyluracil (m 3 U). However, it remains unclear how FTO variously recognizes and catalyzes these diverse substrates. Here we demonstrate-in vitro and in vivo-that FTO has extensive demethylation enzymatic activity on both internal m 6 A and cap m 6 A m Considering that 6mA, m 6 A, and m 6 A m all share the same nucleobase, we present a crystal structure of human FTO bound to 6mA-modified ssDNA, revealing the molecular basis of the catalytic demethylation of FTO toward multiple RNA substrates. We discovered that ( i ) N 6 -methyladenine is the most favorable nucleobase substrate of FTO, ( ii ) FTO displays the same demethylation activity toward internal m 6 A and m 6 A m in the same RNA sequence, suggesting that the substrate specificity of FTO primarily results from the interaction of residues in the catalytic pocket with the nucleobase (rather than the ribose ring), and ( iii ) the sequence and the tertiary structure of RNA can affect the catalytic activity of FTO. Our findings provide a structural basis for understanding the catalytic mechanism through which FTO demethylates its multiple substrates and pave the way forward for the structure-guided design of selective chemicals for functional studies and potential therapeutic applications.
- Synthetic and Functional Biomolecules Center, Beijing National Laboratory for Molecular Sciences, Key Laboratory of Bioorganic Chemistry and Molecular Engineering of Ministry of Education, College of Chemistry and Molecular Engineering, Peking University, Beijing 100871, China.
Organizational Affiliation: