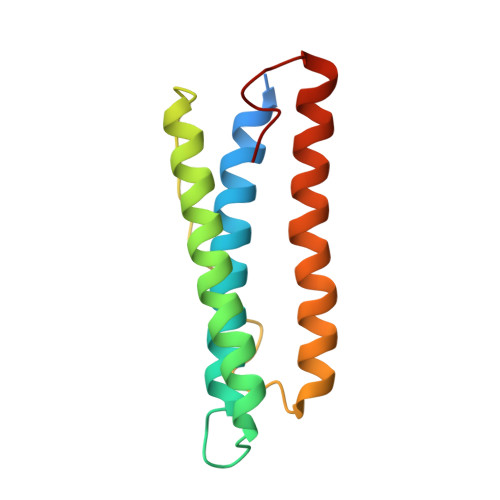
Selective Elimination of the Key Subunit Interfaces Facilitates Conversion of Native 24-mer Protein Nanocage into 8-mer Nanorings.
Wang, W., Wang, L., Chen, H., Zang, J., Zhao, X., Zhao, G., Wang, H.(2018) J Am Chem Soc 140: 14078-14081
- PubMed: 30336004
- DOI: https://doi.org/10.1021/jacs.8b09760
- Primary Citation of Related Structures:
5ZND - PubMed Abstract:
Living systems utilize proteins as building blocks to construct a large variety of self-assembled nanoscale architectures. Yet, creating protein-based assemblies with specific geometries in the laboratory remains challenging. Here, we present a new approach that completely eliminates one natural intersubunit interface of multisubunit protein architecture with high symmetry, resulting in reassembly of the protein architecture into one with lower symmetry. We have applied this approach to the conversion of the 24-mer cage-like ferritin into non-native 8-mer protein nanorings in solution. In the crystal structure, such newly formed nanorings connect with each other through hydrogen bonding in a repeating head-to-tail pattern to form nanotubes, and adjacent nanotubes are staggered relative to one another to create three-dimensional porous protein assemblies. The above strategy allows the study of conversion between protein architectures with different geometries by adjusting the interactions at the intersubunit interfaces, and the fabrication of novel bio-nanomaterials with different geometries.
- Key Laboratory of Chemical Biology and Molecular Engineering of Education Ministry, Institute of Molecular Science , Shanxi University , Taiyuan 030006 , China.
Organizational Affiliation: