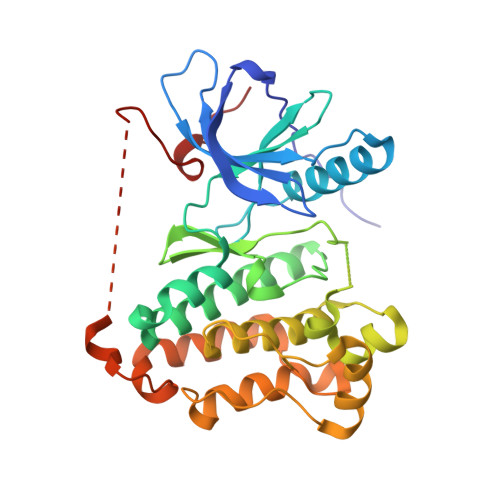
Insight into the Inhibition of Drug-Resistant Mutants of the Receptor Tyrosine Kinase EGFR.
Engel, J., Becker, C., Lategahn, J., Keul, M., Ketzer, J., Muhlenberg, T., Kollipara, L., Schultz-Fademrecht, C., Zahedi, R.P., Bauer, S., Rauh, D.(2016) Angew Chem Int Ed Engl 55: 10909-10912
- PubMed: 27496389
- DOI: https://doi.org/10.1002/anie.201605011
- Primary Citation Related Structures:
5J9Y, 5J9Z - PubMed Abstract:
Targeting acquired drug resistance represents the major challenge in the treatment of EGFR-driven non-small-cell lung cancer (NSCLC). Herein, we describe the structure-based design, synthesis, and biological evaluation of a novel class of covalent EGFR inhibitors that exhibit excellent inhibition of EGFR-mutant drug-resistant cells. Protein X-ray crystallography combined with detailed kinetic studies led to a deeper understanding of the mode of inhibition of EGFR-T790M and provided insight into the key principles for effective inhibition of the recently discovered tertiary mutation at EGFR-C797S.
- Technische Universität Dortmund, Fakultät für Chemie und Chemische Biologie, Otto-Hahn-Strasse 4a, 44227, Dortmund, Germany.
Organizational Affiliation: