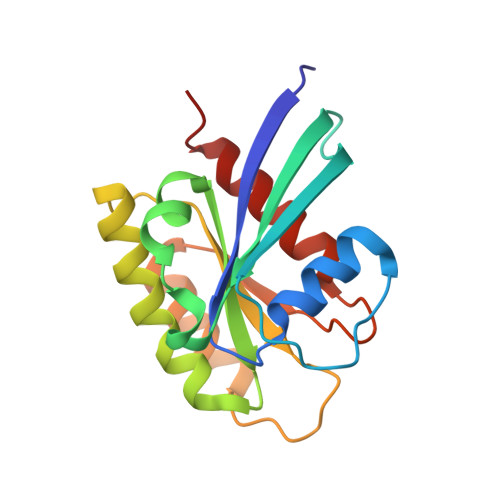
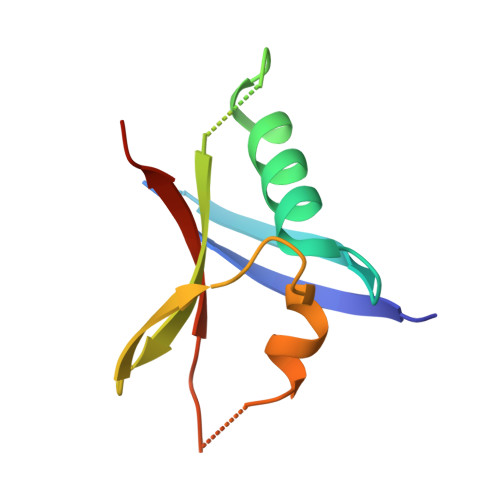
Evolution of AF6-RAS association and its implications in mixed-lineage leukemia.
Smith, M.J., Ottoni, E., Ishiyama, N., Goudreault, M., Haman, A., Meyer, C., Tucholska, M., Gasmi-Seabrook, G., Menezes, S., Laister, R.C., Minden, M.D., Marschalek, R., Gingras, A.C., Hoang, T., Ikura, M.(2017) Nat Commun 8: 1099-1099
- PubMed: 29062045 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-01326-5
- Primary Citation Related Structures:
6AMB - PubMed Abstract:
Elucidation of activation mechanisms governing protein fusions is essential for therapeutic development. MLL undergoes rearrangement with numerous partners, including a recurrent translocation fusing the epigenetic regulator to a cytoplasmic RAS effector, AF6/afadin. We show here that AF6 employs a non-canonical, evolutionarily conserved α-helix to bind RAS, unique to AF6 and the classical RASSF effectors. Further, all patients with MLL-AF6 translocations express fusion proteins missing only this helix from AF6, resulting in exposure of hydrophobic residues that induce dimerization. We provide evidence that oligomerization is the dominant mechanism driving oncogenesis from rare MLL translocation partners and employ our mechanistic understanding of MLL-AF6 to examine how dimers induce leukemia. Proteomic data resolve association of dimerized MLL with gene expression modulators, and inhibiting dimerization disrupts formation of these complexes while completely abrogating leukemogenesis in mice. Oncogenic gene translocations are thus selected under pressure from protein structure/function, underscoring the complex nature of chromosomal rearrangements.
- Institute for Research in Immunology and Cancer, Université de Montréal, Montréal, QC, H3T 1J4, Canada. matthew.james.smith@umontreal.ca.
Organizational Affiliation: