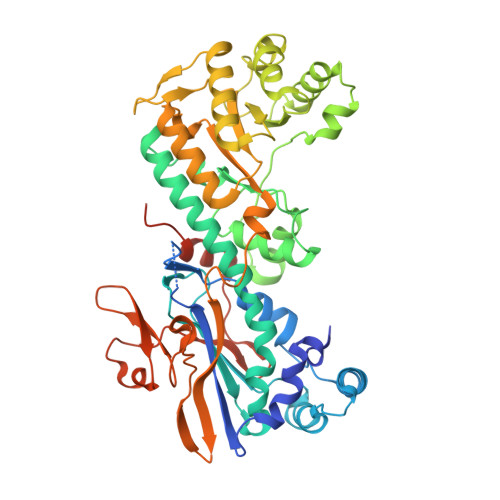
Identification and structure based design of cellularly active cyclo-propyl carboxamide Nicotinamide phosphoribosyltransferase (NAMPT) inhibitors
Palacios, D., Meredith, E., Kawanami, T., Adams, C., Chen, X., Darsigny, V., Geno, E., Palermo, M., Guy, C., Hewett, J., Tierney, L., THigale, S., Weihofen, W.A., Agrikar, U., Boynton, G., George, E., Wang, L.To be published.