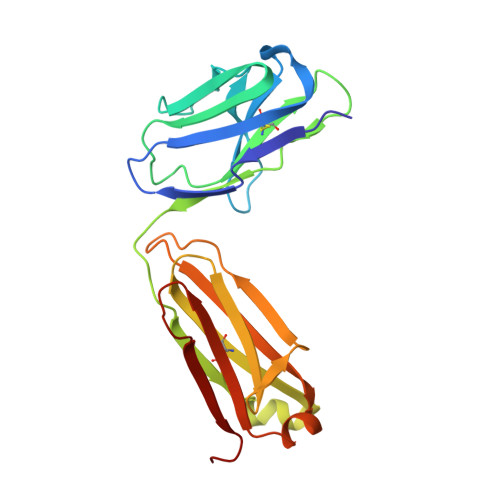
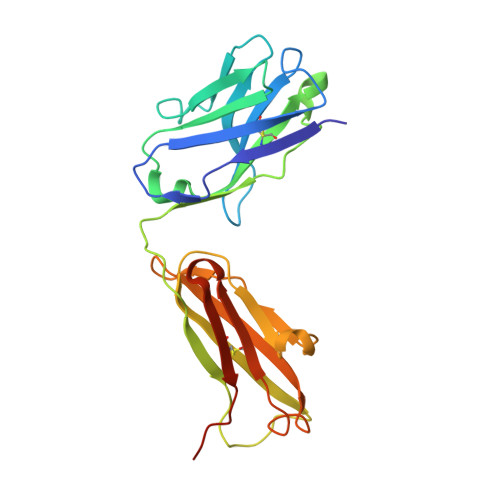
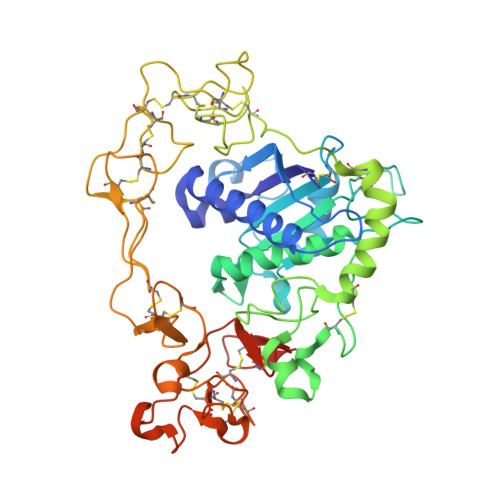
Structural Basis for Regulated Proteolysis by the alpha-Secretase ADAM10.
Seegar, T.C.M., Killingsworth, L.B., Saha, N., Meyer, P.A., Patra, D., Zimmerman, B., Janes, P.W., Rubinstein, E., Nikolov, D.B., Skiniotis, G., Kruse, A.C., Blacklow, S.C.(2017) Cell 171: 1638-1648.e7
- PubMed: 29224781 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2017.11.014
- Primary Citation Related Structures:
6BDZ, 6BE6 - PubMed Abstract:
Cleavage of membrane-anchored proteins by ADAM (a disintegrin and metalloproteinase) endopeptidases plays a key role in a wide variety of biological signal transduction and protein turnover processes. Among ADAM family members, ADAM10 stands out as particularly important because it is both responsible for regulated proteolysis of Notch receptors and catalyzes the non-amyloidogenic α-secretase cleavage of the Alzheimer's precursor protein (APP). We present here the X-ray crystal structure of the ADAM10 ectodomain, which, together with biochemical and cellular studies, reveals how access to the enzyme active site is regulated. The enzyme adopts an unanticipated architecture in which the C-terminal cysteine-rich domain partially occludes the enzyme active site, preventing unfettered substrate access. Binding of a modulatory antibody to the cysteine-rich domain liberates the catalytic domain from autoinhibition, enhancing enzymatic activity toward a peptide substrate. Together, these studies reveal a mechanism for regulation of ADAM activity and offer a roadmap for its modulation.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: