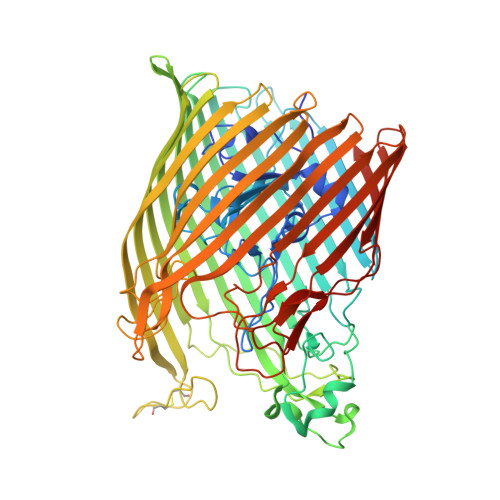
The structure of the bacterial iron-catecholate transporter Fiu suggests that it imports substrates via a two-step mechanism.
Grinter, R., Lithgow, T.(2019) J Biological Chem
- PubMed: 31712312 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.011018
- Primary Citation Related Structures:
6BPM, 6BPN, 6BPO - PubMed Abstract:
The ferric iron uptake (Fiu) transporter from Escherichia coli functions in the transport of iron-catecholate complexes across the bacterial outer membrane, providing the bacterium with iron, which is essential for growth. Recently it has become clear that Fiu also represents a liability for E. coli because its activity allows import of antimicrobial compounds that mimic catecholate. This inadvertent import suggests the potential utility of antimicrobial catechol siderophore mimetics in managing bacterial infections. However, to fully exploit these compounds, a detailed understanding of the mechanism of transport through Fiu and related transporters is required. To address this question, we determined the crystal structure of Fiu at 2.1-2.9 Å and analyzed its function in E. coli Through analysis of the Fiuo crystal structure, in combination with in silico docking and mutagenesis, we provide insight into how Fiu and related transporters bind catecholate in a surface-exposed cavity. Moreover, through determination of the structure of Fiu in multiple crystal states, we revealed the presence of a large, selectively gated cavity in the interior of this transporter. This chamber is large enough to accommodate the Fiu substrate and may allow import of substrates via a two-step mechanism. This would avoid channel formation through the transporter and inadvertent import of toxic molecules. As Fiu and its homologs are the targets of substrate-mimicking antibiotics, these results may assist in the development of these compounds.
- School of Biological Sciences, Monash University, Clayton, 3800 Victoria, Australia rhys.grinter@monash.edu.
Organizational Affiliation: