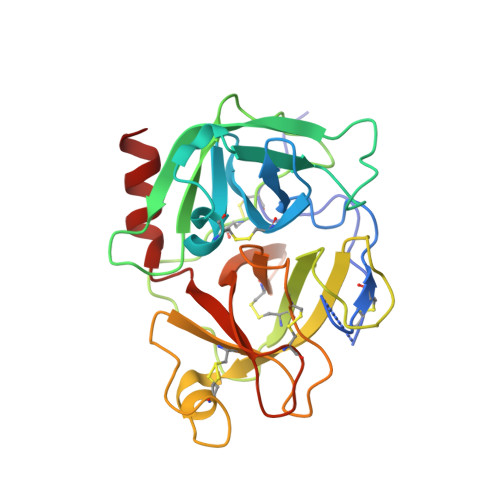
Highly Potent and Selective Plasmin Inhibitors Based on the Sunflower Trypsin Inhibitor-1 Scaffold Attenuate Fibrinolysis in Plasma.
Swedberg, J.E., Wu, G., Mahatmanto, T., Durek, T., Caradoc-Davies, T.T., Whisstock, J.C., Law, R.H.P., Craik, D.J.(2019) J Med Chem 62: 552-560
- PubMed: 30520638 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01139
- Primary Citation Related Structures:
6D3X, 6D3Y, 6D3Z, 6D40 - PubMed Abstract:
Antifibrinolytic drugs provide important pharmacological interventions to reduce morbidity and mortality from excessive bleeding during surgery and after trauma. Current drugs used for inhibiting the dissolution of fibrin, the main structural component of blood clots, are associated with adverse events due to lack of potency, high doses, and nonselective inhibition mechanisms. These drawbacks warrant the development of a new generation of highly potent and selective fibrinolysis inhibitors. Here, we use the 14-amino acid backbone-cyclic sunflower trypsin inhibitor-1 scaffold to design a highly potent ( K i = 0.05 nM) inhibitor of the primary serine protease in fibrinolysis, plasmin. This compound displays a million-fold selectivity over other serine proteases in blood, inhibits fibrinolysis in plasma more effectively than the gold-standard therapeutic inhibitor aprotinin, and is a promising candidate for development of highly specific fibrinolysis inhibitors with reduced side effects.
- Institute for Molecular Bioscience , The University of Queensland , Brisbane , QLD 4072 , Australia.
Organizational Affiliation: