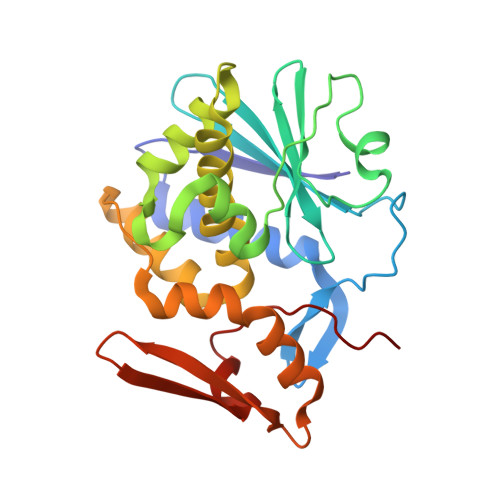
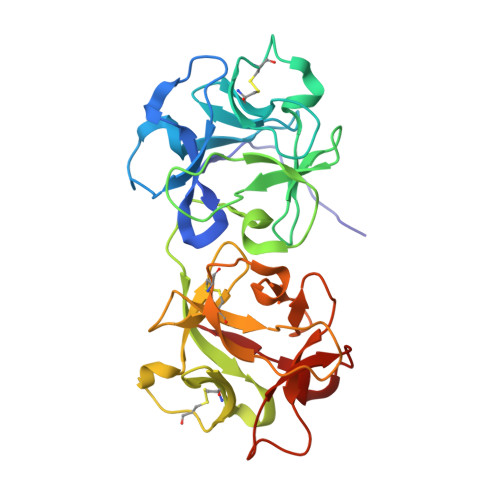
Crystal Structure of Mistletoe Lectin I (ML-I) from Viscum album in Complex with 4-N-Furfurylcytosine at 2.85 angstrom Resolution.
Ahmad, M.S., Rasheed, S., Falke, S., Khaliq, B., Perbandt, M., Choudhary, M.I., Markiewicz, W.T., Barciszewski, J., Betzel, C.(2018) Med Chem 14: 754-763
- PubMed: 29792147
- DOI: https://doi.org/10.2174/1573406414666180524095946
- Primary Citation of Related Structures:
6ELY - PubMed Abstract:
Viscum album (the European mistletoe) is a semi-parasitic plant, which is of high medical interest. It is widely found in Europe, Asia, and North America. It contains at least three distinct lectins (i.e. ML-I, II, and III), varying in molecular mass and specificity. Among them, ML-I is in focus of medical research for various activities, including anti-cancer activities. To understand the molecular basis for such medical applications, a few studies have already addressed the structural and functional analysis of ML-I in complex with ligands. In continuation of these efforts, we are reporting the crystal structure of ML from Viscum album in complex with the nucleic acid oxidation product 4-N-furfurylcytosine (FC) refined to 2.85 Å resolution. FC is known to be involved in different metabolic pathways related to oxidative stress and DNA modification. X-ray suitable hexagonal crystals of the ML-I/FC complex were grown within four days at 294 K using the hanging drop vapor diffusion method. Diffraction data were collected up to a resolution of 2.85 Å. The ligand affinity was verified by in-silico docking. The high-resolution structure was refined subsequently to analyze particularly the active site conformation and a binding epitope of 4-N-furfurylcytosine. A distinct 2Fo-Fc electron density at the active site was interpreted as a single FC molecule. The specific binding of FC is achieved also through hydrophobic interactions involving Tyr76A, Tyr115A, Glu165A, and Leu157A of the ML-I A-chain. The binding energy of FC to the active site of ML-I was calculated as well to be -6.03 kcal mol-1. In comparison to other reported ML-I complexes, we observed distinct differences in the vicinity of the nucleic acid base binding site upon interaction with FC. Therefore, data obtained will provide new insights in understanding the specificity, inhibition, and cytotoxicity of the ML-I A-chain, and related RIPs.
- Laboratory for Structural Biology of Infection and Inflammation, University of Hamburg, c/o DESY, Build. 22a, Notkestrasse 85, 22607 Hamburg, Germany.
Organizational Affiliation: