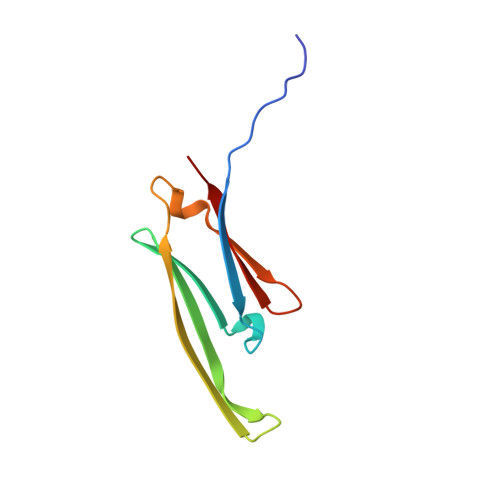
HspB1 phosphorylation regulates its intramolecular dynamics and mechanosensitive molecular chaperone interaction with filamin C.
Collier, M.P., Alderson, T.R., de Villiers, C.P., Nicholls, D., Gastall, H.Y., Allison, T.M., Degiacomi, M.T., Jiang, H., Mlynek, G., Furst, D.O., van der Ven, P.F.M., Djinovic-Carugo, K., Baldwin, A.J., Watkins, H., Gehmlich, K., Benesch, J.L.P.(2019) Sci Adv 5: eaav8421-eaav8421
- PubMed: 31131323 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aav8421
- Primary Citation Related Structures:
6GJH - PubMed Abstract:
Mechanical force-induced conformational changes in proteins underpin a variety of physiological functions, typified in muscle contractile machinery. Mutations in the actin-binding protein filamin C (FLNC) are linked to musculoskeletal pathologies characterized by altered biomechanical properties and sometimes aggregates. HspB1, an abundant molecular chaperone, is prevalent in striated muscle where it is phosphorylated in response to cues including mechanical stress. We report the interaction and up-regulation of both proteins in three mouse models of biomechanical stress, with HspB1 being phosphorylated and FLNC being localized to load-bearing sites. We show how phosphorylation leads to increased exposure of the residues surrounding the HspB1 phosphosite, facilitating their binding to a compact multidomain region of FLNC proposed to have mechanosensing functions. Steered unfolding of FLNC reveals that its extension trajectory is modulated by the phosphorylated region of HspB1. This may represent a posttranslationally regulated chaperone-client protection mechanism targeting over-extension during mechanical stress.
- Department of Chemistry, Physical and Theoretical Chemistry Laboratory, University of Oxford, South Parks Road, Oxford OX1 3QZ, UK.
Organizational Affiliation: