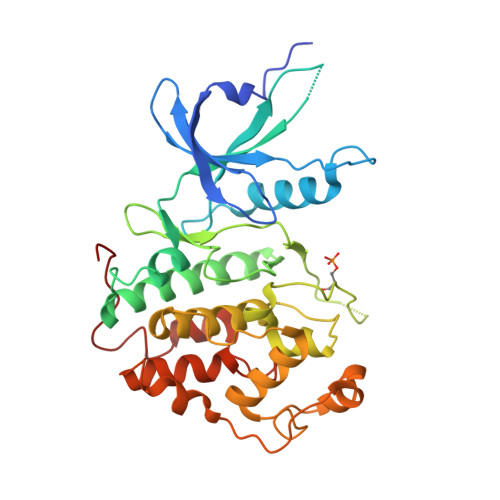
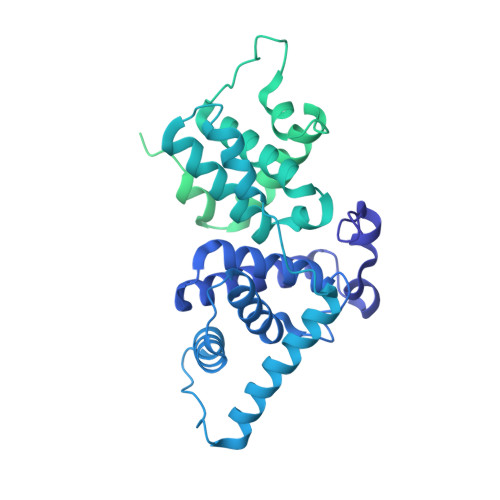
Small Molecules Co-targeting CKI alpha and the Transcriptional Kinases CDK7/9 Control AML in Preclinical Models.
Minzel, W., Venkatachalam, A., Fink, A., Hung, E., Brachya, G., Burstain, I., Shaham, M., Rivlin, A., Omer, I., Zinger, A., Elias, S., Winter, E., Erdman, P.E., Sullivan, R.W., Fung, L., Mercurio, F., Li, D., Vacca, J., Kaushansky, N., Shlush, L., Oren, M., Levine, R., Pikarsky, E., Snir-Alkalay, I., Ben-Neriah, Y.(2018) Cell 175: 171-185.e25
- PubMed: 30146162 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2018.07.045
- Primary Citation Related Structures:
6GZD, 6GZH, 6GZM - PubMed Abstract:
CKIα ablation induces p53 activation, and CKIα degradation underlies the therapeutic effect of lenalidomide in a pre-leukemia syndrome. Here we describe the development of CKIα inhibitors, which co-target the transcriptional kinases CDK7 and CDK9, thereby augmenting CKIα-induced p53 activation and its anti-leukemic activity. Oncogene-driving super-enhancers (SEs) are highly sensitive to CDK7/9 inhibition. We identified multiple newly gained SEs in primary mouse acute myeloid leukemia (AML) cells and demonstrate that the inhibitors abolish many SEs and preferentially suppress the transcription elongation of SE-driven oncogenes. We show that blocking CKIα together with CDK7 and/or CDK9 synergistically stabilize p53, deprive leukemia cells of survival and proliferation-maintaining SE-driven oncogenes, and induce apoptosis. Leukemia progenitors are selectively eliminated by the inhibitors, explaining their therapeutic efficacy with preserved hematopoiesis and leukemia cure potential; they eradicate leukemia in MLL-AF9 and Tet2 -/- ;Flt3 ITD AML mouse models and in several patient-derived AML xenograft models, supporting their potential efficacy in curing human leukemia.
- The Lautenberg Center for Immunology and Cancer Research, Institute of Medical Research Israel-Canada, Hebrew University-Hadassah Medical School, Jerusalem, Israel.
Organizational Affiliation: