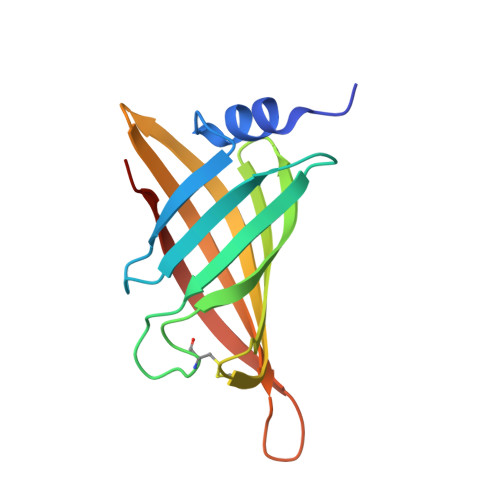
Crystal structure of afifavidin reveals common features of molecular assemblage in the bacterial dimeric avidins.
Avraham, O., Bayer, E.A., Livnah, O.(2018) FEBS J 285: 4617-4630
- PubMed: 30369031 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14685
- Primary Citation Related Structures:
6HDS, 6HDT, 6HDV - PubMed Abstract:
The subfamily of bacterial dimeric avidins is being extended through the discovery of additional members originating from diverse sources. All of these newly discovered dimeric avidin forms exhibit high affinity towards biotin, despite their lack of critical Trp in the classical tetrameric forms. The common feature of forming cylinder-like multimers (hexamers and octamers) seems to be more than a random occurrence, which generally characterizes their apo forms in the crystalline state and also in some cases in solution. Afifavidin from the Gram-negative α-proteobacterium Afifella pfennigii is the fourth member of the subfamily of dimers, which, in the intact apo form, also congregates into octamers both in the solution and in the crystalline state, whereby the C-terminal extended segments stretch into the biotin-binding sites of adjacent non-canonical monomers. The intact apo afifavidin molecule self-assembles into toroid-shaped nanostructures that dissociate into the inherent dimers upon binding biotin. On removal of the C-terminal regions, the short-form of afifavidin forms dimers both in the solution and in the crystalline states. The high affinity of the dimeric forms of afifavidin towards biotin is maintained, due to the conserved disulfide bridge between L3,4 and L5,6 and the presence of Phe50 in L3,4 that compensate for the lack of the critical Trp in the tetrameric avidins. These cyclic multimeric-avidin assemblies may be exploited in the future to further diversify biotin-based nanotechnology or to serve as building blocks in the construction of bio-inspired materials. DATABASE: Structural data are available in the PDB databases under the accession numbers: 6HDV, 6HDS, 6HDT.
- Department of Biological Chemistry, The Alexander Silverman Institute of Life Sciences, The Wolfson Centre for Applied Structural Biology, The Hebrew University of Jerusalem, Israel.
Organizational Affiliation: