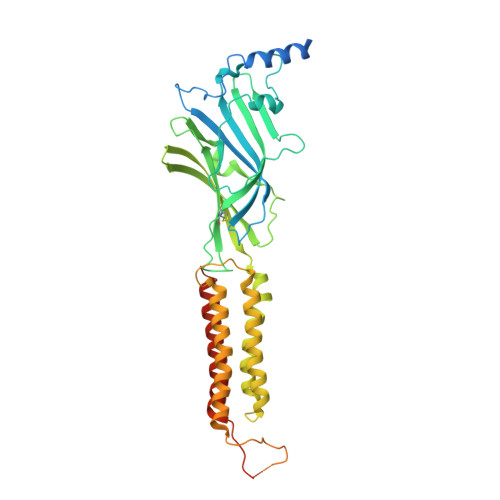
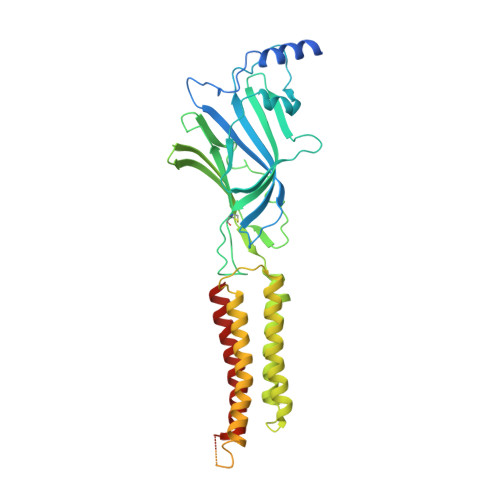
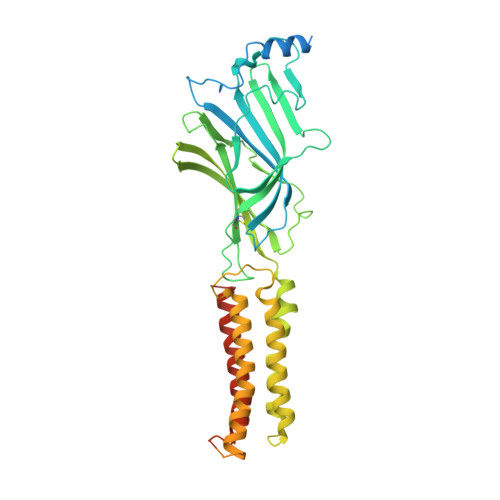
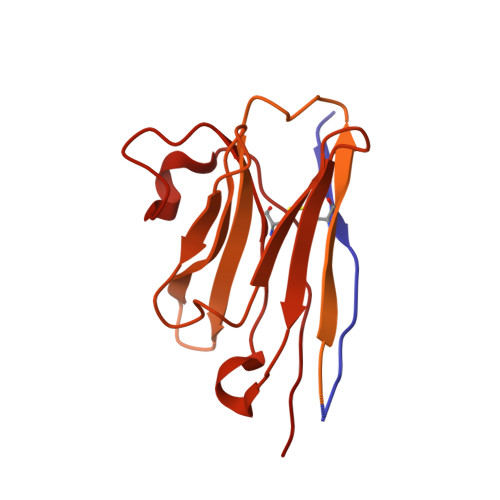
GABAAreceptor signalling mechanisms revealed by structural pharmacology.
Masiulis, S., Desai, R., Uchanski, T., Serna Martin, I., Laverty, D., Karia, D., Malinauskas, T., Zivanov, J., Pardon, E., Kotecha, A., Steyaert, J., Miller, K.W., Aricescu, A.R.(2019) Nature 565: 454-459
- PubMed: 30602790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0832-5
- Primary Citation Related Structures:
6HUG, 6HUJ, 6HUK, 6HUO, 6HUP - PubMed Abstract:
Type-A γ-aminobutyric (GABA A ) receptors are ligand-gated chloride channels with a very rich pharmacology. Some of their modulators, including benzodiazepines and general anaesthetics, are among the most successful drugs in clinical use and are common substances of abuse. Without reliable structural data, the mechanistic basis for the pharmacological modulation of GABA A receptors remains largely unknown. Here we report several high-resolution cryo-electron microscopy structures in which the full-length human α1β3γ2L GABA A receptor in lipid nanodiscs is bound to the channel-blocker picrotoxin, the competitive antagonist bicuculline, the agonist GABA (γ-aminobutyric acid), and the classical benzodiazepines alprazolam and diazepam. We describe the binding modes and mechanistic effects of these ligands, the closed and desensitized states of the GABA A receptor gating cycle, and the basis for allosteric coupling between the extracellular, agonist-binding region and the transmembrane, pore-forming region. This work provides a structural framework in which to integrate previous physiology and pharmacology research and a rational basis for the development of GABA A receptor modulators.
- MRC Laboratory of Molecular Biology, Cambridge Biomedical Campus, Cambridge, UK. simonasm@mrc-lmb.cam.ac.uk.
Organizational Affiliation: