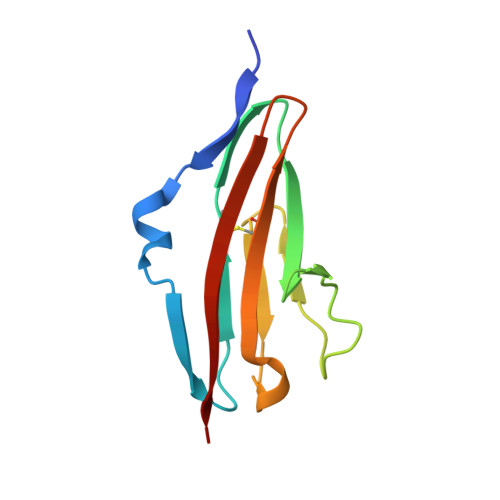
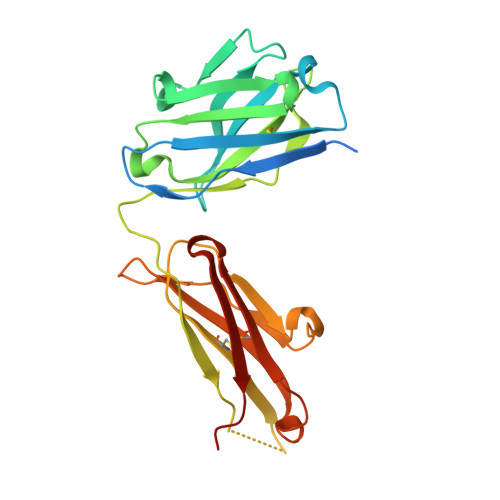
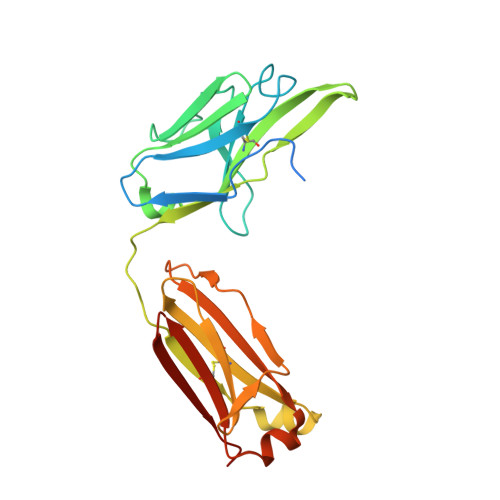
Preclinical Development of U3-1784, a Novel FGFR4 Antibody Against Cancer, and Avoidance of Its On-target Toxicity.
Bartz, R., Fukuchi, K., Ohtsuka, T., Lange, T., Gruner, K., Watanabe, I., Hayashi, S., Oda, Y., Kawaida, R., Komori, H., Kashimoto, Y., Wirtz, P., Mayer, J.A., Redondo-Muller, M., Saito, S., Takahashi, M., Hanzawa, H., Imai, E., Martinez, A., Hanai, M., Haussinger, D., Chapman, R.W., Agatsuma, T., Bange, J., Abraham, R.(2019) Mol Cancer Ther 18: 1832-1843
- PubMed: 31350344 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-18-0048
- Primary Citation Related Structures:
6J6Y - PubMed Abstract:
The FGFR4/FGF19 signaling axis is overactivated in 20% of liver tumors and currently represents a promising targetable signaling mechanism in this cancer type. However, blocking FGFR4 or FGF19 has proven challenging due to its physiological role in suppressing bile acid synthesis which leads to increased toxic bile acid plasma levels upon FGFR4 inhibition. An FGFR4-targeting antibody, U3-1784, was generated in order to investigate its suitability as a cancer treatment without major side effects.U3-1784 is a high-affinity fully human antibody that was obtained by phage display technology and specifically binds to FGFR4. The antibody inhibits cell signaling by competing with various FGFs for their FGFR4 binding site thereby inhibiting receptor activation and downstream signaling via FRS2 and Erk. The inhibitory effect on tumor growth was investigated in 10 different liver cancer models in vivo The antibody specifically slowed tumor growth of models overexpressing FGF19 by up to 90% whereas tumor growth of models not expressing FGF19 was unaffected. In cynomolgus monkeys, intravenous injection of U3-1784 caused elevated serum bile acid and liver enzyme levels indicating potential liver damage. These effects could be completely prevented by the concomitant oral treatment with the bile acid sequestrant colestyramine, which binds and eliminates bile acids in the gut. These results offer a new biomarker-driven treatment modality in liver cancer without toxicity and they suggest a general strategy for avoiding adverse events with FGFR4 inhibitors.
- U3 Pharma GmbH/Daiichi-Sankyo, Martinsried, Germany.
Organizational Affiliation: