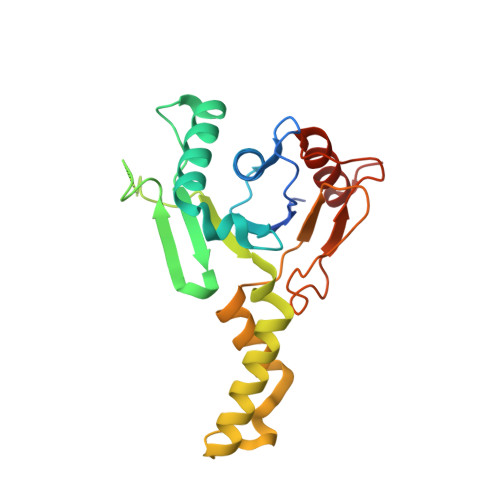
Structural insights to heterodimeric cis-prenyltransferases through yeast dehydrodolichyl diphosphate synthase subunit Nus1.
Ma, J., Ko, T.P., Yu, X., Zhang, L., Ma, L., Zhai, C., Guo, R.T., Liu, W., Li, H., Chen, C.C.(2019) Biochem Biophys Res Commun 515: 621-626
- PubMed: 31178134 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.05.135
- Primary Citation Related Structures:
6JCN - PubMed Abstract:
The polyprenoid glycan carriers are produced by cis-prenyltransferases (cis-PTs), which function as heterodimers in metazoa and fungi or homodimers in bacteria, but both are found in plants, protista and archaea. Heterodimeric cis-PTs comprise catalytic and non-catalytic subunits while homodimeric enzymes contain two catalytic subunits. The non-catalytic subunits of cis-PT shows low sequence similarity to known cis-PTs and their structure information is of great interests. Here we report the crystal structure of Nus1, the non-catalytic subunit of cis-PT from Saccharomyces cerevisiae. We also investigate the heterodimer formation and active site conformation by constructing a homology model of Nus1 and its catalytic subunit. Nus1 does not contain an active site, but its C-terminus may participate in catalysis by interacting with the substrates bound to the catalytic subunit. These results provide important basis for further investigation of heterodimeric cis-PTs.
- Key Laboratory of Industrial Biotechnology, Ministry of Education, School of Biotechnology, Jiangnan University, Wuxi, Jiangsu, 214122, China; State Key Laboratory of Biocatalysis and Enzyme Engineering, Hubei Collaborative Innovation Center for Green Transformation of Bio-Resources, Hubei Key Laboratory of Industrial Biotechnology, School of Life Sciences, Hubei University, Wuhan, 430062, China.
Organizational Affiliation: