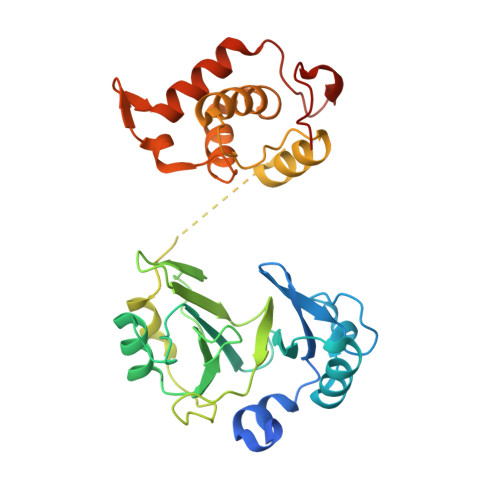
Crystal structures of phage NrS-1 N300-dNTPs-Mg2+complex provide molecular mechanisms for substrate specificity.
Guo, H., Li, M., Wu, H., Wang, W., Yu, F., He, J.(2019) Biochem Biophys Res Commun 515: 551-557
- PubMed: 31176489 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.05.162
- Primary Citation Related Structures:
6JON, 6JOP, 6JOQ - PubMed Abstract:
A novel DNA polymerase from the deep-sea vent phage NrS-1, was characterized as a primase-polymerase (referred to as prim-pol), which works as a self-priming DNA polymerase to synthesize de novo long DNA strands. Functional research on the NrS-1 prim-pol illustrated that the N-terminal 300 residues (referred to as N300) have de novo synthesis activity similar to that of the full-length enzyme. Just like other prim-pols, NrS-1 prim-pol was able to initiate DNA synthesis, proficiently discriminating against ribonucleotides (NTPs), exclusively using deoxynucleotides (dNTPs). However, the structural basis for this discrimination is not well understood. Here, the three kinds of crystal structures of N300-dNTPs-Mg 2+ complex were determined. These complex structures shared the identical steric architecture and hydrogen-bond interactions in the catalytic center. The results of biochemical studies indicated that R145 possibly plays an indispensable role in the primer extension. Mutagenesis and structural simulation showed that the backbone carboxyl group of Y146, as a potential sugar selector, was involved in steric clashing with the incoming 2'-OH group of NTPs. However, the mechanism of substrate discrimination probably was different from that of other prim-pols, according to the structural analyses and sequence comparison.
- Shanghai Institute of Applied Physics, Chinese Academy of Sciences, Shanghai, 201800, China; University of Chinese Academy of Sciences, Beijing, 100049, China.
Organizational Affiliation: