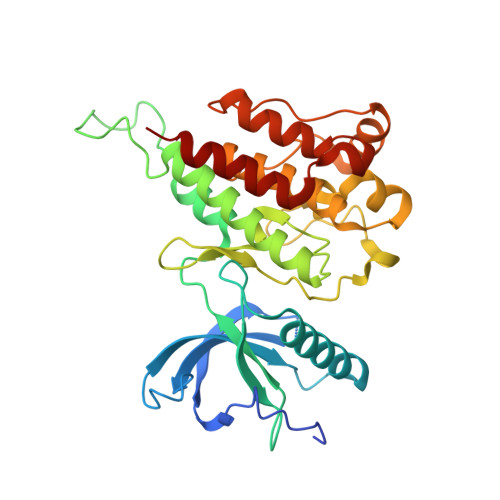
Development of a Potent and Specific FGFR4 Inhibitor for the Treatment of Hepatocellular Carcinoma.
Rezende Miranda, R., Fu, Y., Chen, X., Perino, J., Cao, P., Carpten, J., Chen, Y., Zhang, C.(2020) J Med Chem 63: 11484-11497
- PubMed: 33030342 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.0c00044
- Primary Citation Related Structures:
6JPE - PubMed Abstract:
Abnormal activation of the fibroblast growth factor 19 (FGF19)/fibroblast growth factor receptor 4 (FGFR4) signaling pathway has been shown to drive the proliferation of a significant portion of hepatocellular carcinoma (HCC). Resistance and toxicity are serious drawbacks that have been observed upon use of the current first- and second-line treatment options for HCC, therefore warranting the investigation of alternative therapeutic approaches. We report the development and biological characterization of a covalent inhibitor that is highly potent and exquisitely specific to FGFR4. The crystal structure of this inhibitor in complex with FGFR4 was solved, confirming its covalent binding and revealing its binding mode. We also describe the first clickable probe for FGFR4 that can be used to directly measure target engagement in cells. Our compound exhibited great antitumor activity in HCC cell lines and tumor xenograft models. These results provide evidence of a promising therapeutic lead for the treatment of a subset of HCC patients.
- Department of Chemistry and Loker Hydrocarbon Research Institute, University of Southern California, Los Angeles, California 90089, United States.
Organizational Affiliation: