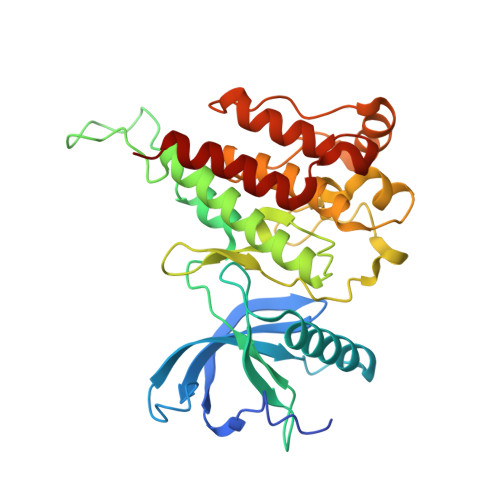
Characterization of FGF401 as a reversible covalent inhibitor of fibroblast growth factor receptor 4.
Zhou, Z., Chen, X., Fu, Y., Zhang, Y., Dai, S., Li, J., Chen, L., Xu, G., Chen, Z., Chen, Y.(2019) Chem Commun (Camb) 55: 5890-5893
- PubMed: 31041948 Search on PubMed
- DOI: https://doi.org/10.1039/c9cc02052g
- Primary Citation Related Structures:
6JPJ - PubMed Abstract:
Biochemical and structural studies provide information on the mode of action of FGF401 as a selective, reversible covalent inhibitor of FGFR4. Kinase and proliferation assays reveal that FGF401 has the ability to overcome gatekeeper mutations in FGFR4.
- NHC Key Laboratory of Cancer Proteomics & Laboratory of Structural Biology, Xiangya Hospital, Central South University, Changsha, Hunan 410008, China. yonghenc@163.com.
Organizational Affiliation: