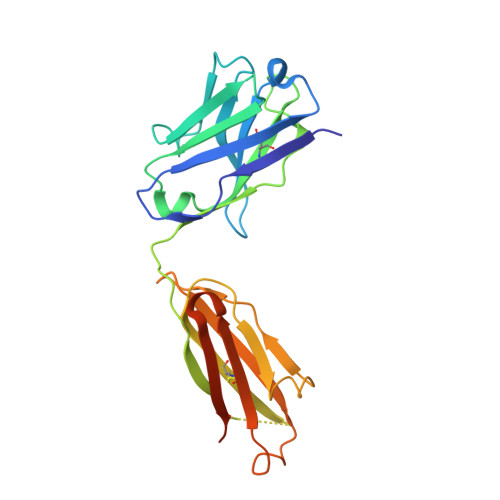
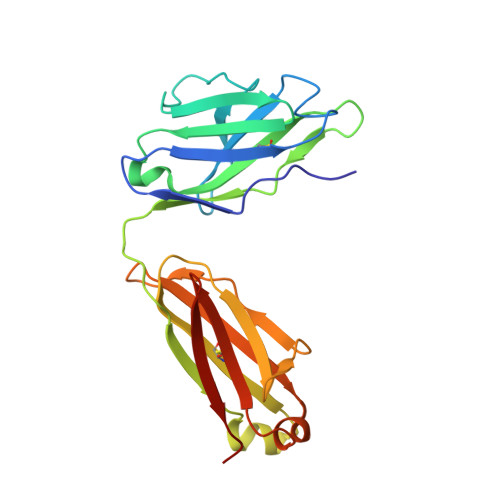
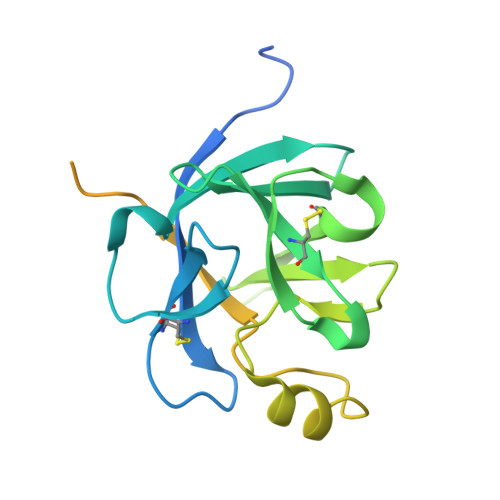
Novel Abs targeting the N-terminus of fibroblast growth factor 19 inhibit hepatocellular carcinoma growth without bile-acid-related side-effects.
Liu, H., Zheng, S., Hou, X., Liu, X., Du, K., Lv, X., Li, Y., Yang, F., Li, W., Sui, J.(2020) Cancer Sci 111: 1750-1760
- PubMed: 32061104 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/cas.14353
- Primary Citation Related Structures:
6KTR - PubMed Abstract:
Hepatocellular carcinoma (HCC) is a common and particularly fatal form of cancer for which very few drugs are effective. The fibroblast growth factor 19 (FGF19) has been viewed as a driver of HCC development and a potential Ab target for developing novel HCC therapy. However, a previously developed anti-FGF19 Ab disrupted FGF19's normal regulatory function and caused severe bile-acid-related side-effects despite of having potent antitumor effects in preclinical models. Here, we developed novel human Abs (G1A8 and HS29) that specifically target the N-terminus of FGF19. Both Abs inhibited FGF19-induced HCC cell proliferation in vitro and significantly suppressed HCC tumor growth in mouse models. Importantly, no bile-acid-related side effects were observed in preclinical cynomolgus monkeys. Fundamentally, our study demonstrates that it is possible to target FGF19 for anti-HCC therapies without adversely affecting its normal bile acid regulatory function, and highlights the exciting promise of G1A8 or HS29 as potential therapy for HCC.
- National Institute of Biological Sciences (NIBS), Beijing, China.
Organizational Affiliation: