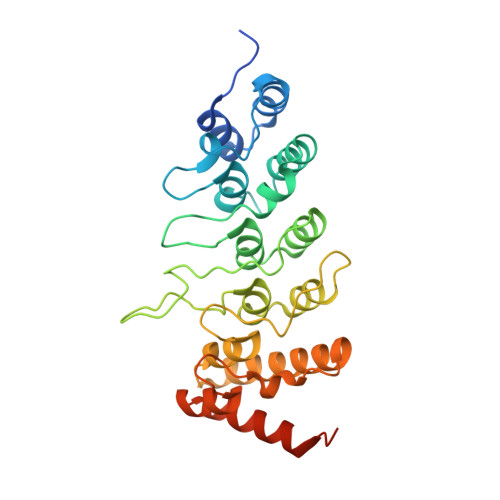
Structure determination of the human TRPV1 ankyrin-repeat domain under nonreducing conditions.
Tanaka, M., Hayakawa, K., Ogawa, N., Kurokawa, T., Kitanishi, K., Ite, K., Matsui, T., Mori, Y., Unno, M.(2020) Acta Crystallogr F Struct Biol Commun 76: 130-137
- PubMed: 32133998 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20001533
- Primary Citation Related Structures:
6L93 - PubMed Abstract:
TRPV1, a member of the transient receptor potential (TRP) channels family, has been found to be involved in redox sensing. The crystal structure of the human TRPV1 ankyrin-repeat domain (TRPV1-ARD) was determined at 4.5 Å resolution under nonreducing conditions. This is the first report of the crystal structure of a ligand-free form of TRPV1-ARD and in particular of the human homologue. The structure showed a unique conformation in finger loop 3 near Cys258, which is most likely to be involved in inter-subunit disulfide-bond formation. Also, in human TRPV1-ARD it was possible for solvent to access Cys258. This structural feature might be related to the high sensitivity of human TRPV1 to oxidants. ESI-MS revealed that Cys258 did not form an S-OH functionality even under nonreducing conditions.
- Graduate School of Science and Engineering, Ibaraki University, Hitachi, Ibaraki 316-8511, Japan.
Organizational Affiliation: