Structural insights into secretory immunoglobulin A and its interaction with a pneumococcal adhesin.
Wang, Y., Wang, G., Li, Y., Zhu, Q., Shen, H., Gao, N., Xiao, J.(2020) Cell Res 30: 602-609
- PubMed: 32398862 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-020-0336-3
- Primary Citation Related Structures:
6LX3, 6LXW - PubMed Abstract:
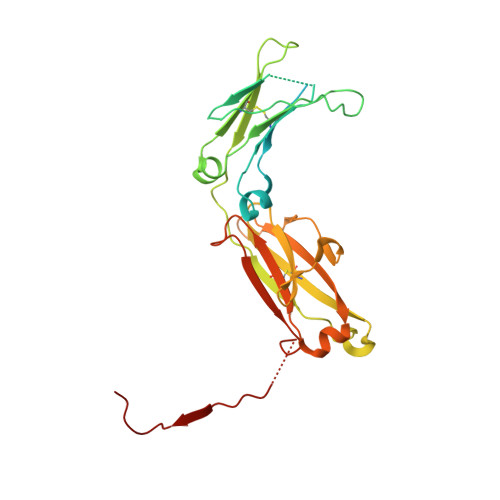
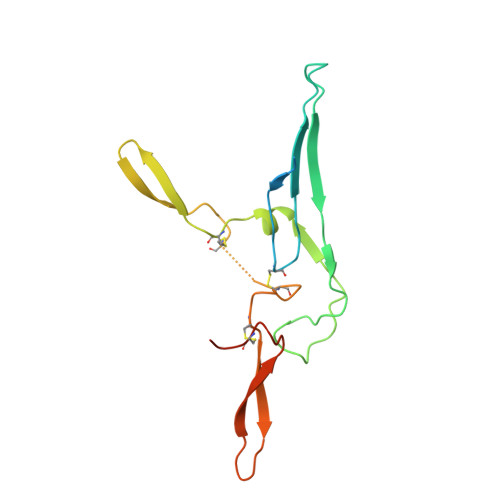
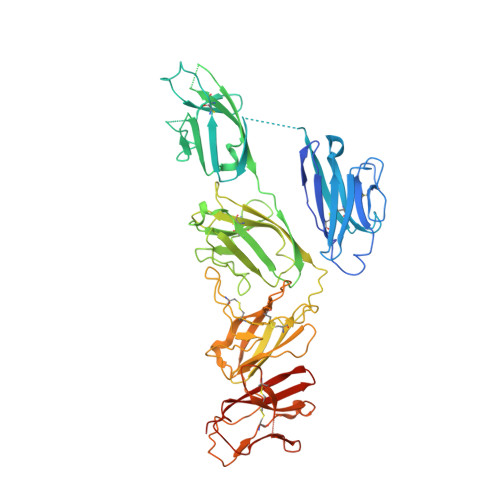
Secretory Immunoglobulin A (SIgA) is the most abundant antibody at the mucosal surface. It possesses two additional subunits besides IgA: the joining chain (J-chain) and secretory component (SC). SC is the ectodomain of the polymeric immunoglobulin receptor (pIgR), which functions to transport IgA to the mucosa. How the J-chain and pIgR/SC facilitate the assembly and secretion of SIgA remains incompletely understood. Furthermore, during the infection of Streptococcus pneumoniae, the pneumococcal adhesin SpsA hijacks pIgR/SC and SIgA to gain entry to human cells and evade host defense. How SpsA targets pIgR/SC and SIgA also remains elusive. Here we report a cryo-electron microscopy structure of the Fc region of IgA1 (Fcα) in complex with the J-chain and SC (Fcα-J-SC), which reveals the organization principle of SIgA. We also present a structure of Fcα-J-SC complexed with SpsA, which uncovers the specific interactions between SpsA and human pIgR/SC. These results advance the molecular understanding of SIgA and shed light on S. pneumoniae pathogenesis.
- State Key Laboratory of Protein and Plant Gene Research, Peking University, Beijing, 100871, China.
Organizational Affiliation: