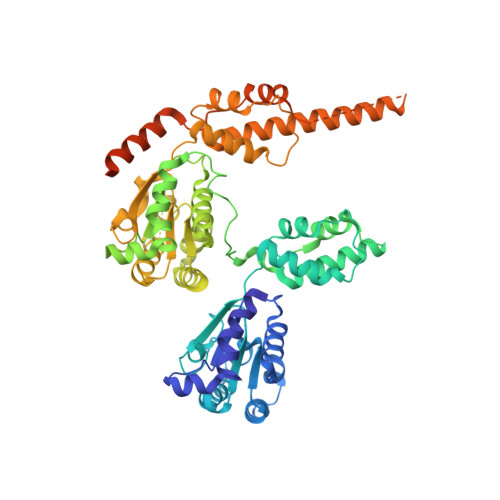
Structural Basis of p97 Inhibition by the Site-Selective Anticancer Compound CB-5083.
Tang, W.K., Odzorig, T., Jin, W., Xia, D.(2019) Mol Pharmacol 95: 286-293
- PubMed: 30591537 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1124/mol.118.114256
- Primary Citation Related Structures:
6MCK - PubMed Abstract:
Inhibition of p97, a key player in the ubiquitin-proteasome degradation pathway, has been proposed as a treatment of cancer. This concept was nearly realized recently when a potent p97 inhibitor, 1-[4-(benzylamino)-5H,7H,8H-pyrano[4,3-d]pyrimidin-2-yl]-2-methyl-1H-indole-4-carboxamide (CB-5083), was developed and demonstrated broad antitumor activity in various tumor models. CB-5083 functions as a competitive inhibitor that binds selectively to the ATP-binding site of the D2 domain, although both the D1 and D2 ATPase sites of p97 are highly similar. Despite its promising anticancer activity, CB-5083 failed its phase I clinical trials due to an unexpected off-target effect, which necessitates further improvement of the inhibitor. In this study, we determined the crystal structure of N-terminal domain-truncated p97 in complex with CB-5083. It provides a structural basis for the specificity of CB-5083 toward the D2 domain, offers an explanation in atomic detail for the mutations that confer resistance to CB-5083, and establishes a foundation for future structure-guided efforts to develop the next generation of p97 inhibitors.
- Laboratory of Cell Biology, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, Maryland.
Organizational Affiliation: