A mutually induced conformational fit underlies Ca2+-directed interactions between calmodulin and the proximal C terminus of KCNQ4 K+channels.
Archer, C.R., Enslow, B.T., Taylor, A.B., De la Rosa, V., Bhattacharya, A., Shapiro, M.S.(2019) J Biological Chem 294: 6094-6112
- PubMed: 30808708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.006857
- Primary Citation Related Structures:
6N5W - PubMed Abstract:
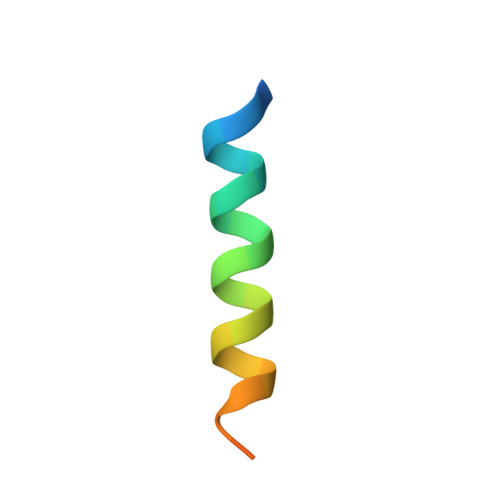
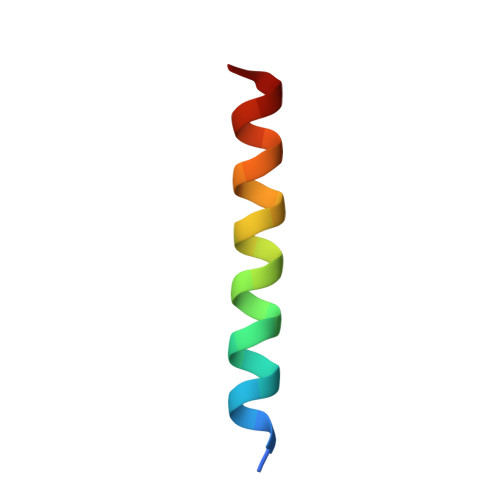
Calmodulin (CaM) conveys intracellular Ca 2+ signals to KCNQ (Kv7, "M-type") K + channels and many other ion channels. Whether this "calmodulation" involves a dramatic structural rearrangement or only slight perturbations of the CaM/KCNQ complex is as yet unclear. A consensus structural model of conformational shifts occurring between low nanomolar and physiologically high intracellular [Ca 2+ ] is still under debate. Here, we used various techniques of biophysical chemical analyses to investigate the interactions between CaM and synthetic peptides corresponding to the A and B domains of the KCNQ4 subtype. We found that in the absence of CaM, the peptides are disordered, whereas Ca 2+ /CaM imposed helical structure on both KCNQ A and B domains. Isothermal titration calorimetry revealed that Ca 2+ /CaM has higher affinity for the B domain than for the A domain of KCNQ2-4 and much higher affinity for the B domain when prebound with the A domain. X-ray crystallography confirmed that these discrete peptides spontaneously form a complex with Ca 2+ /CaM, similar to previous reports of CaM binding KCNQ-AB domains that are linked together. Microscale thermophoresis and heteronuclear single-quantum coherence NMR spectroscopy indicated the C-lobe of Ca 2+ -free CaM to interact with the KCNQ4 B domain ( K d ∼10-20 μm), with increasing Ca 2+ molar ratios shifting the CaM-B domain interactions via only the CaM C-lobe to also include the N-lobe. Our findings suggest that in response to increased Ca 2+ , CaM undergoes lobe switching that imposes a dramatic mutually induced conformational fit to both the proximal C terminus of KCNQ4 channels and CaM, likely underlying Ca 2+ -dependent regulation of KCNQ gating.
- From the Departments of Cell and Integrative Physiology, University of Texas Health San Antonio, San Antonio, Texas 78229; Departments of Biochemistry and Structural Biology, University of Texas Health San Antonio, San Antonio, Texas 78229.
Organizational Affiliation: