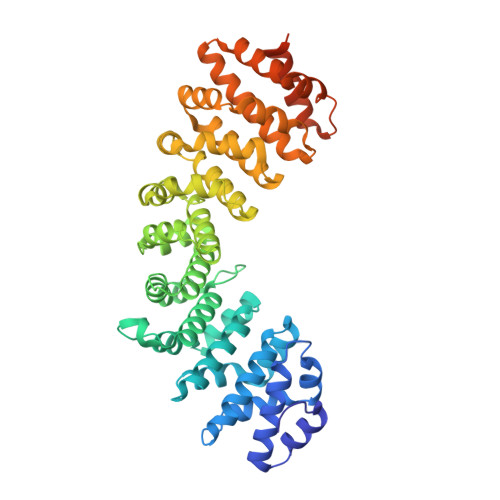
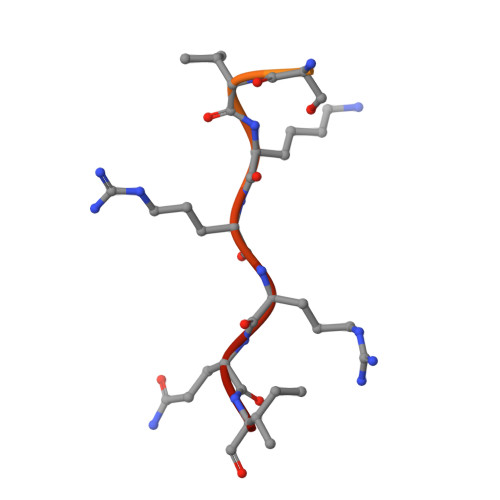
Comparative study of the interactions between fungal transcription factor nuclear localization sequences with mammalian and fungal importin-alpha.
Bernardes, N.E., Fukuda, C.A., da Silva, T.D., de Oliveira, H.C., de Barros, A.C., Dreyer, T.R., Bertolini, M.C., Fontes, M.R.M.(2020) Sci Rep 10: 1458-1458
- PubMed: 31996719
- DOI: https://doi.org/10.1038/s41598-020-58316-9
- Primary Citation of Related Structures:
6P6A, 6P6E - PubMed Abstract:
Importin-α (Impα) is an adaptor protein that binds to cargo proteins (containing Nuclear Localization Sequences - NLSs), for their translocation to the nucleus. The specificities of the Impα/NLS interactions have been studied, since these features could be used as important tools to find potential NLSs in nuclear proteins or even for the development of targets to inhibit nuclear import or to design peptides for drug delivery. Few structural studies have compared different Impα variants from the same organism or Impα of different organisms. Previously, we investigated nuclear transport of transcription factors with Neurospora crassa Impα (NcImpα). Herein, NIT-2 and PAC-3 transcription factors NLSs were studied in complex with Mus musculus Impα (MmImpα). Calorimetric assays demonstrated that the PAC-3 NLS peptide interacts with both Impα proteins with approximately the same affinity. The NIT-2 NLS sequence binds with high affinity to the Impα major binding site from both organisms, but its binding to minor binding sites reveals interesting differences due to the presence of additional interactions of NIT-2-NLS with MmImpα. These findings, together with previous results with Impα from other organisms, indicate that the differential affinity of NLSs to minor binding sites may be also responsible for the selectivity of some cargo proteins recognition and transport.
- Departamento de Física e Biofísica, Instituto de Biociências, Universidade Estadual Paulista (UNESP), Botucatu, São Paulo, Brazil.
Organizational Affiliation: