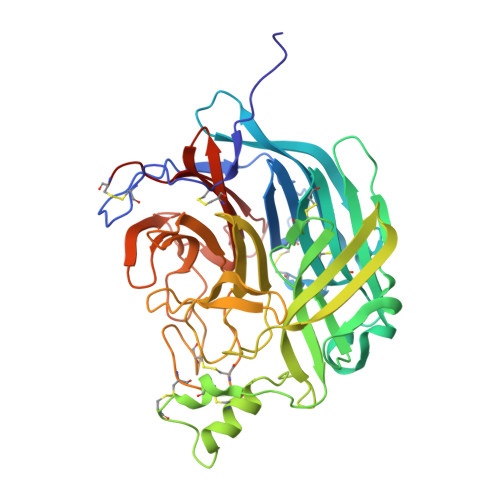
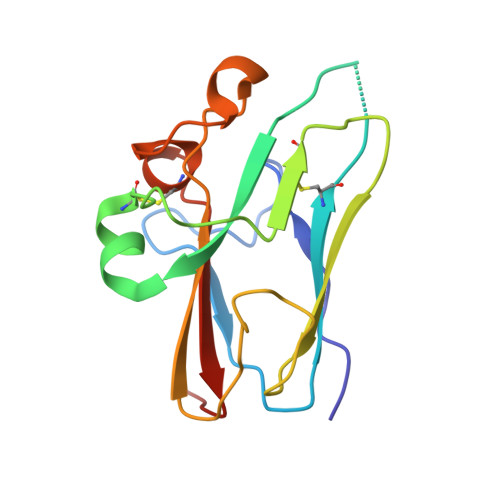
Structural and functional analyses reveal promiscuous and species specific use of ephrin receptors by Cedar virus.
Laing, E.D., Navaratnarajah, C.K., Cheliout Da Silva, S., Petzing, S.R., Xu, Y., Sterling, S.L., Marsh, G.A., Wang, L.F., Amaya, M., Nikolov, D.B., Cattaneo, R., Broder, C.C., Xu, K.(2019) Proc Natl Acad Sci U S A 116: 20707-20715
- PubMed: 31548390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1911773116
- Primary Citation Related Structures:
6P72, 6P7S, 6P7Y - PubMed Abstract:
Cedar virus (CedV) is a bat-borne henipavirus related to Nipah virus (NiV) and Hendra virus (HeV), zoonotic agents of fatal human disease. CedV receptor-binding protein (G) shares only ∼30% sequence identity with those of NiV and HeV, although they can all use ephrin-B2 as an entry receptor. We demonstrate that CedV also enters cells through additional B- and A-class ephrins (ephrin-B1, ephrin-A2, and ephrin-A5) and report the crystal structure of the CedV G ectodomain alone and in complex with ephrin-B1 or ephrin-B2. The CedV G receptor-binding site is structurally distinct from other henipaviruses, underlying its capability to accommodate additional ephrin receptors. We also show that CedV can enter cells through mouse ephrin-A1 but not human ephrin-A1, which differ by 1 residue in the key contact region. This is evidence of species specific ephrin receptor usage by a henipavirus, and implicates additional ephrin receptors in potential zoonotic transmission.
- Department of Microbiology and Immunology, Uniformed Services University of the Health Sciences, Bethesda, MD 20814.
Organizational Affiliation: