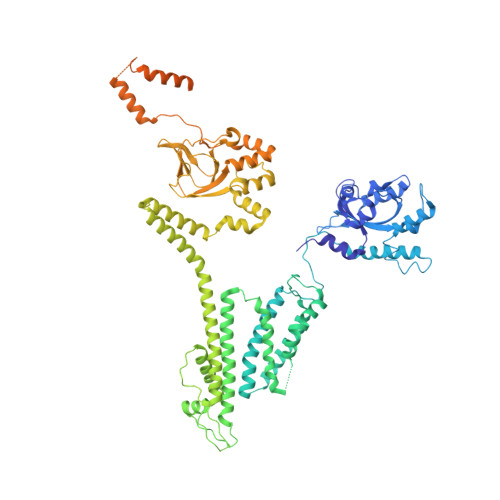
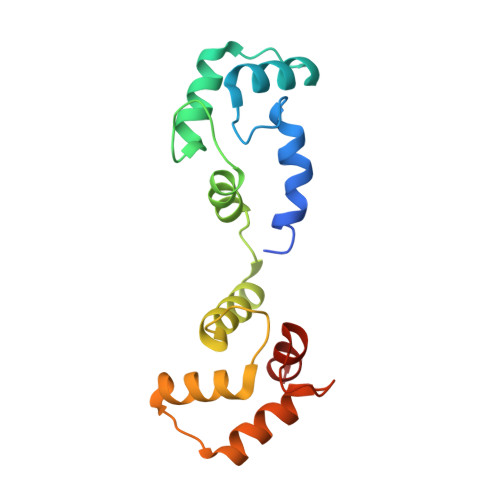
Regulation of Eag1 gating by its intracellular domains.
Whicher, J.R., MacKinnon, R.(2019) Elife 8
- PubMed: 31490124 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.49188
- Primary Citation Related Structures:
6PBX, 6PBY - PubMed Abstract:
Voltage-gated potassium channels (K v s) are gated by transmembrane voltage sensors (VS) that move in response to changes in membrane voltage. K v 10.1 or Eag1 also has three intracellular domains: PAS, C-linker, and CNBHD. We demonstrate that the Eag1 intracellular domains are not required for voltage-dependent gating but likely interact with the VS to modulate gating. We identified specific interactions between the PAS, CNBHD, and VS that modulate voltage-dependent gating and provide evidence that VS movement destabilizes these interactions to promote channel opening. Additionally, mutation of these interactions renders Eag1 insensitive to calmodulin inhibition. The structure of the calmodulin insensitive mutant in a pre-open conformation suggests that channel opening may occur through a rotation of the intracellular domains and calmodulin may prevent this rotation by stabilizing interactions between the VS and intracellular domains. Intracellular domains likely play a similar modulatory role in voltage-dependent gating of the related K v 11-12 channels.
- Laboratory of Molecular Neurobiology and Biophysics, The Rockefeller University, Howard Hughes Medical Institute, New York, United States.
Organizational Affiliation: