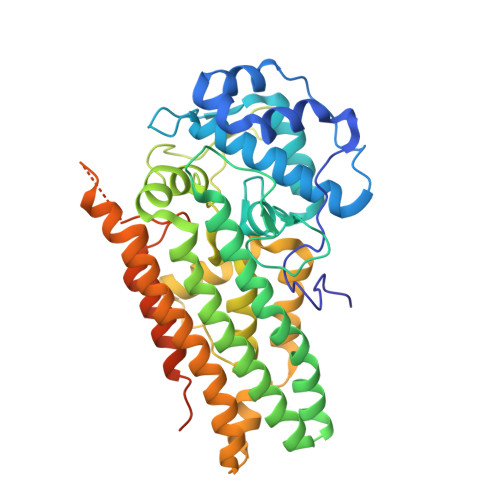
Structural Basis of Inhibitor Selectivity in Human Indoleamine 2,3-Dioxygenase 1 and Tryptophan Dioxygenase.
Pham, K.N., Lewis-Ballester, A., Yeh, S.R.(2019) J Am Chem Soc 141: 18771-18779
- PubMed: 31682426 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.9b08871
- Primary Citation Related Structures:
6PYY, 6PYZ, 6PZ1 - PubMed Abstract:
Indoleamine 2,3-dioxygenase 1 (hIDO1) and tryptophan dioxygenase (hTDO) are two of the only three heme-based dioxygenases in humans. They have recently been identified as key cancer immunotherapeutic drug targets. While structures of hIDO1 in complex with inhibitors have been documented, so far there are no structures of hTDO-inhibitor complexes available. Here we use PF-06840003 (IPD), a hIDO1-selective inhibitor in clinical trials, as a structural probe to elucidate inhibitor-selectivity in hIDO1 versus hTDO. Spectroscopic studies show that IPD exhibits 400-fold higher inhibition activity toward hIDO1 with respect to hTDO. Crystallographic structures reveal that the binding pocket of IPD in the active site in hIDO1 is much more flexible as compared to that in hTDO, which offers a molecular explanation for the superior inhibition activity of IPD in hIDO1 with respect to hTDO. In addition to the IPD bound in the active site, a second IPD molecule was identified in an inhibitory site on the proximal side of the heme in hIDO1 and in an exosite that is ∼40 Å away from the active site in hTDO. Taken together the data provide new insights into structure-based design of mono and dual inhibitors targeting hIDO1 and/or hTDO.
- Department of Physiology and Biophysics , Albert Einstein College of Medicine , The Bronx , New York 10461 , United States.
Organizational Affiliation: