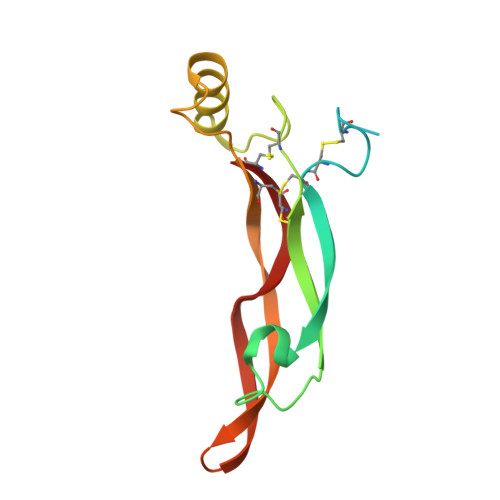
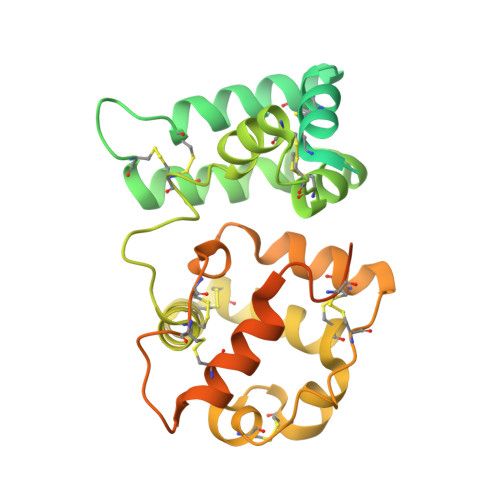
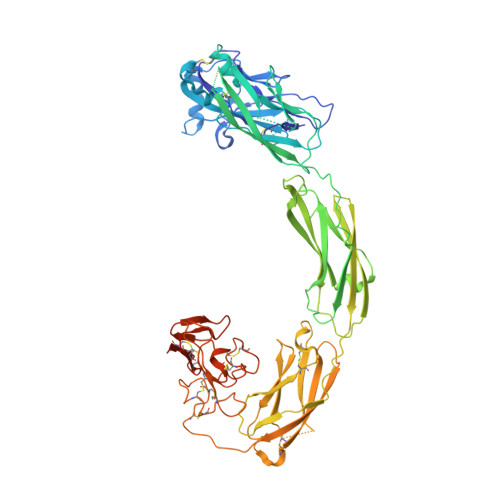
Cryo-EM analyses reveal the common mechanism and diversification in the activation of RET by different ligands.
Li, J., Shang, G., Chen, Y.J., Brautigam, C.A., Liou, J., Zhang, X., Bai, X.C.(2019) Elife 8
- PubMed: 31535977 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.47650
- Primary Citation Related Structures:
6Q2J, 6Q2N, 6Q2O, 6Q2R, 6Q2S - PubMed Abstract:
RET is a receptor tyrosine kinase (RTK) that plays essential roles in development and has been implicated in several human diseases. Different from most of RTKs, RET requires not only its cognate ligands but also co-receptors for activation, the mechanisms of which remain unclear due to lack of high-resolution structures of the ligand/co-receptor/receptor complexes. Here, we report cryo-EM structures of the extracellular region ternary complexes of GDF15/GFRAL/RET, GDNF/GFRα1/RET, NRTN/GFRα2/RET and ARTN/GFRα3/RET. These structures reveal that all the four ligand/co-receptor pairs, while using different atomic interactions, induce a specific dimerization mode of RET that is poised to bring the two kinase domains into close proximity for cross-phosphorylation. The NRTN/GFRα2/RET dimeric complex further pack into a tetrameric assembly, which is shown by our cell-based assays to regulate the endocytosis of RET. Our analyses therefore reveal both the common mechanism and diversification in the activation of RET by different ligands.
- Department of Biophysics, University of Texas Southwestern Medical Center, Dallas, United States.
Organizational Affiliation: