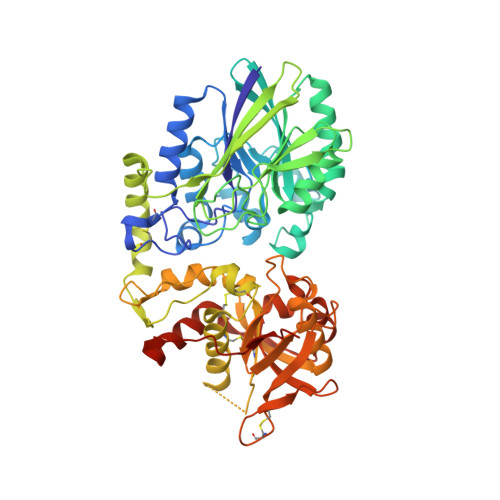
X-Ray Co-Crystal Structure Guides the Way to Subnanomolar Competitive Ecto-5'-Nucleotidase (CD73) Inhibitors for Cancer Immunotherapy
Bhattarai, S., Pippel, J., Meyer, A., Freundlieb, M., Schmies, C., Abdelrahman, A., Fiene, A., Lee, S.Y., Zimmermann, H., El-Tayeb, A., Yegutkin, G.G., Strater, N., Muller, C.E.(2019) Adv Ther