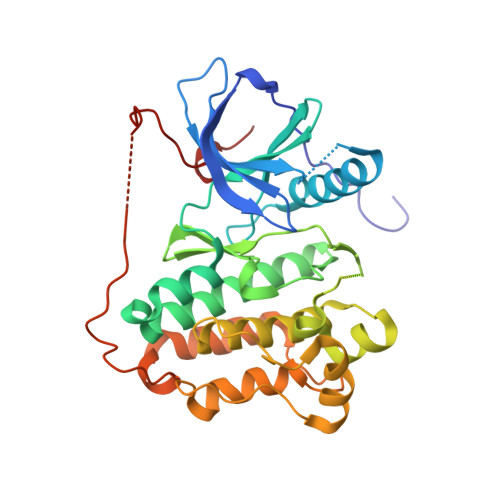
Start Selective and Rigidify: The Discovery Path toward a Next Generation of EGFR Tyrosine Kinase Inhibitors.
Engelhardt, H., Bose, D., Petronczki, M., Scharn, D., Bader, G., Baum, A., Bergner, A., Chong, E., Dobel, S., Egger, G., Engelhardt, C., Ettmayer, P., Fuchs, J.E., Gerstberger, T., Gonnella, N., Grimm, A., Grondal, E., Haddad, N., Hopfgartner, B., Kousek, R., Krawiec, M., Kriz, M., Lamarre, L., Leung, J., Mayer, M., Patel, N.D., Simov, B.P., Reeves, J.T., Schnitzer, R., Schrenk, A., Sharps, B., Solca, F., Stadtmuller, H., Tan, Z., Wunberg, T., Zoephel, A., McConnell, D.B.(2019) J Med Chem 62: 10272-10293
- PubMed: 31689114 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01169
- Primary Citation Related Structures:
6S9B, 6S9C, 6S9D - PubMed Abstract:
The epidermal growth factor receptor (EGFR), when carrying an activating mutation like del19 or L858R, acts as an oncogenic driver in a subset of lung tumors. While tumor responses to tyrosine kinase inhibitors (TKIs) are accompanied by marked tumor shrinkage, the response is usually not durable. Most patients relapse within two years of therapy often due to acquisition of an additional mutation in EGFR kinase domain that confers resistance to TKIs. Crucially, oncogenic EGFR harboring both resistance mutations, T790M and C797S, can no longer be inhibited by currently approved EGFR TKIs. Here, we describe the discovery of BI-4020 , which is a noncovalent, wild-type EGFR sparing, macrocyclic TKI. BI-4020 potently inhibits the above-described EGFR variants and induces tumor regressions in a cross-resistant EGFR del19 T790M C797S xenograft model. Key was the identification of a highly selective but moderately potent benzimidazole followed by complete rigidification of the molecule through macrocyclization.
- Boehringer Ingelheim RCV GmbH & Co KG , Dr-Boehringer-Gasse 5-11 , Vienna 1120 , Austria.
Organizational Affiliation: