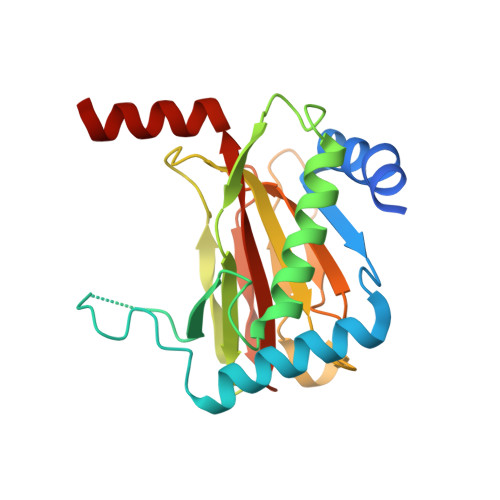
Structure-Activity Relationship and Crystallographic Studies on 4-Hydroxypyrimidine HIF Prolyl Hydroxylase Domain Inhibitors.
Holt-Martyn, J.P., Chowdhury, R., Tumber, A., Yeh, T.L., Abboud, M.I., Lippl, K., Lohans, C.T., Langley, G.W., Figg Jr., W., McDonough, M.A., Pugh, C.W., Ratcliffe, P.J., Schofield, C.J.(2020) ChemMedChem 15: 270-273
- PubMed: 31751494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.201900557
- Primary Citation Related Structures:
6ST3 - PubMed Abstract:
The 2-oxoglutarate-dependent hypoxia inducible factor prolyl hydroxylases (PHDs) are targets for treatment of a variety of diseases including anaemia. One PHD inhibitor is approved for use for the treatment of renal anaemia and others are in late stage clinical trials. The number of reported templates for PHD inhibition is limited. We report structure-activity relationship and crystallographic studies on a promising class of 4-hydroxypyrimidine-containing PHD inhibitors.
- Department of Chemistry, University of Oxford Chemistry Research Laboratory, 12 Mansfield Road, Oxford, OX1 3TA, UK.
Organizational Affiliation: