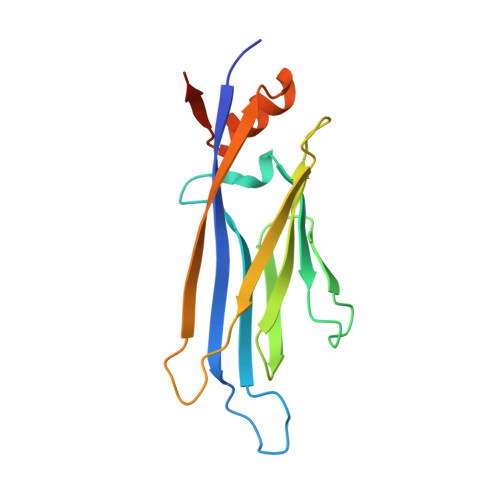
Structural Insights into Interaction Mechanisms of Alternative Piperazine-urea YEATS Domain Binders in MLLT1.
Ni, X., Heidenreich, D., Christott, T., Bennett, J., Moustakim, M., Brennan, P.E., Fedorov, O., Knapp, S., Chaikuad, A.(2019) ACS Med Chem Lett 10: 1661-1666
- PubMed: 31857843 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.9b00460
- Primary Citation Related Structures:
6T1I, 6T1J, 6T1L, 6T1M, 6T1N, 6T1O - PubMed Abstract:
YEATS-domain-containing MLLT1 is an acetyl/acyl-lysine reader domain, which is structurally distinct from well-studied bromodomains and has been strongly associated in development of cancer. Here, we characterized piperazine-urea derivatives as an acetyl/acyl-lysine mimetic moiety for MLLT1. Crystal structures revealed distinct interaction mechanisms of this chemotype compared to the recently described benzimidazole-amide based inhibitors, exploiting different binding pockets within the protein. Thus, the piperazine-urea scaffold offers an alternative strategy for targeting the YEATS domain family.
- Institute of Pharmaceutical Chemistry, Goethe-University Frankfurt, 60438 Frankfurt, Germany.
Organizational Affiliation: