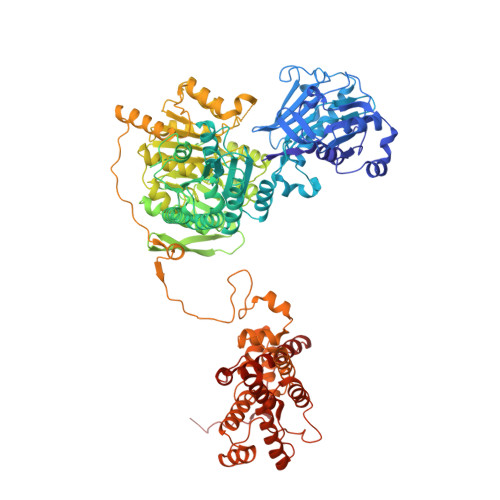
Molecular basis for acetyl-CoA production by ATP-citrate lyase
Wei, X., Schultz, K., Bazilevsky, G.A., Vogt, A., Marmorstein, R.(2020) Nat Struct Mol Biol 27: 33-41
- PubMed: 31873304 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0351-6
- Primary Citation Related Structures:
6POE, 6POF, 6UI9, 6UIA, 6UUW, 6UUZ, 6UV5 - PubMed Abstract:
ATP-citrate lyase (ACLY) synthesizes cytosolic acetyl coenzyme A (acetyl-CoA), a fundamental cellular building block. Accordingly, aberrant ACLY activity is observed in many diseases. Here we report cryo-EM structures of human ACLY, alone or bound to substrates or products. ACLY forms a homotetramer with a rigid citrate synthase homology (CSH) module, flanked by four flexible acetyl-CoA synthetase homology (ASH) domains; CoA is bound at the CSH-ASH interface in mutually exclusive productive or unproductive conformations. The structure of a catalytic mutant of ACLY in the presence of ATP, citrate and CoA substrates reveals a phospho-citryl-CoA intermediate in the ASH domain. ACLY with acetyl-CoA and oxaloacetate products shows the products bound in the ASH domain, with an additional oxaloacetate in the CSH domain, which could function in ACLY autoinhibition. These structures, which are supported by biochemical and biophysical data, challenge previous proposals of the ACLY catalytic mechanism and suggest additional therapeutic possibilities for ACLY-associated metabolic disorders.
- Department of Biochemistry and Biophysics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, USA.
Organizational Affiliation: