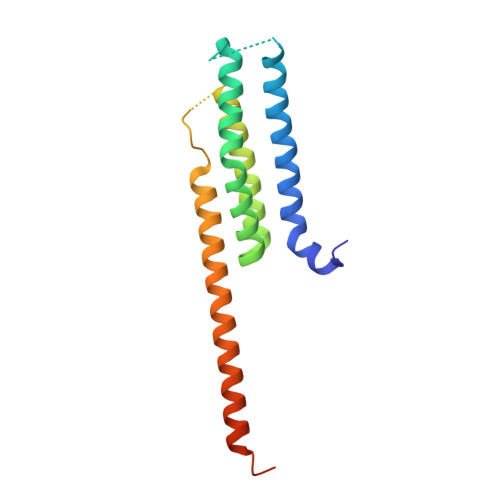
Structure-based, multi-targeted drug discovery approach to eicosanoid inhibition: Dual inhibitors of mPGES-1 and 5-lipoxygenase activating protein (FLAP).
Ho, J.D., Lee, M.R., Rauch, C.T., Aznavour, K., Park, J.S., Luz, J.G., Antonysamy, S., Condon, B., Maletic, M., Zhang, A., Hickey, M.J., Hughes, N.E., Chandrasekhar, S., Sloan, A.V., Gooding, K., Harvey, A., Yu, X.P., Kahl, S.D., Norman, B.H.(2020) Biochim Biophys Acta Gen Subj 1865: 129800-129800
- PubMed: 33246032 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2020.129800
- Primary Citation Related Structures:
6VGC, 6VGI, 6VL4 - PubMed Abstract:
Due to the importance of both prostaglandins (PGs) and leukotrienes (LTs) as pro-inflammatory mediators, and the potential for eicosanoid shunting in the presence of pathway target inhibitors, we have investigated an approach to inhibiting the formation of both PGs and LTs as part of a multi-targeted drug discovery effort. We generated ligand-protein X-ray crystal structures of known inhibitors of microsomal prostaglandin E2 synthase-1 (mPGES-1) and the 5-Lipoxygenase Activating Protein (FLAP), with their respective proteins, to understand the overlapping pharmacophores. We subsequently used molecular modeling and structure-based drug design (SBDD) to identify hybrid structures intended to inhibit both targets. This work enabled the preparation of compounds 4 and 5, which showed potent in vitro inhibition of both targets. Our findings enhance the structural understanding of mPGES-1 and FLAP's unique ligand binding pockets and should accelerate the discovery of additional dual inhibitors for these two important integral membrane protein drug targets.
- Lilly Biotechnology Center, San Diego, CA 92121, USA. Electronic address: ho_joseph_d@lilly.com.
Organizational Affiliation: