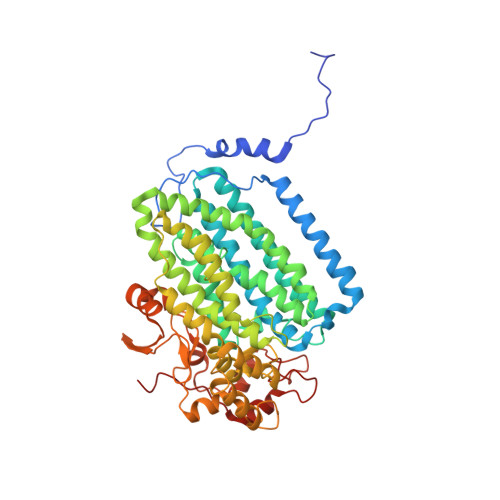
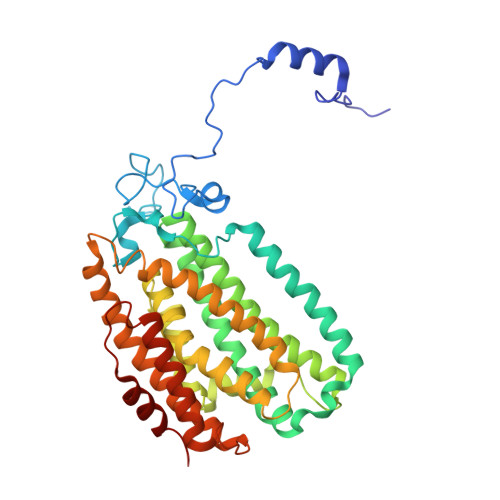
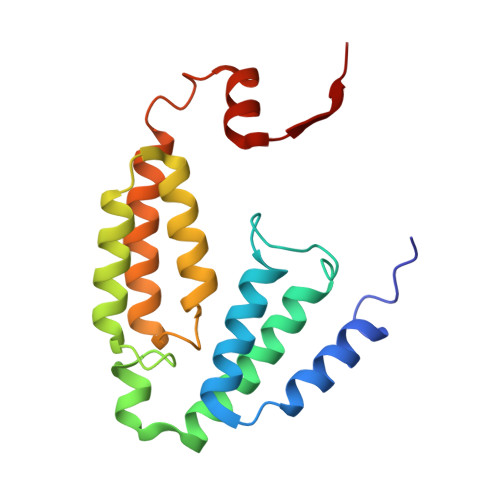
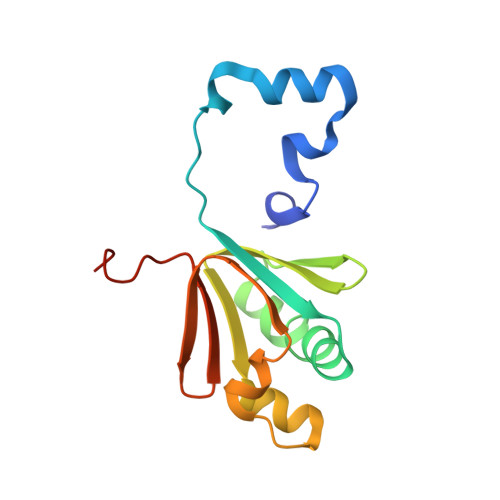
Structural Studies of theMethylosinus trichosporiumOB3b Soluble Methane Monooxygenase Hydroxylase and Regulatory Component Complex Reveal a Transient Substrate Tunnel.
Jones, J.C., Banerjee, R., Shi, K., Aihara, H., Lipscomb, J.D.(2020) Biochemistry 59: 2946-2961
- PubMed: 32692178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.0c00459
- Primary Citation Related Structures:
6VK4, 6VK5, 6VK6, 6VK7, 6VK8 - PubMed Abstract:
The metalloenzyme soluble methane monooxygenase (sMMO) consists of hydroxylase (sMMOH), regulatory (MMOB), and reductase components. When sMMOH forms a complex with MMOB, the rate constants are greatly increased for the sequential access of O 2 , protons, and CH 4 to an oxygen-bridged diferrous metal cluster located in the buried active site. Here, we report high-resolution X-ray crystal structures of the diferric and diferrous states of both sMMOH and the sMMOH:MMOB complex using the components from Methylosinus trichosporium OB3b. These structures are analyzed for O 2 access routes enhanced when the complex forms. Previously reported, lower-resolution structures of the sMMOH:MMOB complex from the sMMO of Methylococcus capsulatus Bath revealed a series of cavities through sMMOH postulated to serve as the O 2 conduit. This potential role is evaluated in greater detail using the current structures. Additionally, a search for other potential O 2 conduits in the M. trichosporium OB3b sMMOH:MMOB complex revealed a narrow molecular tunnel, termed the W308-tunnel. This tunnel is sized appropriately for O 2 and traverses the sMMOH-MMOB interface before accessing the active site. The kinetics of reaction of O 2 with the diferrous sMMOH:MMOB complex in solution show that use of the MMOB V41R variant decreases the rate constant for O 2 binding >25000-fold without altering the component affinity. The location of Val41 near the entrance to the W308-tunnel is consistent with the tunnel serving as the primary route for the transfer of O 2 into the active site. Accordingly, the crystal structures show that formation of the diferrous sMMOH:MMOB complex restricts access through the chain of cavities while opening the W308-tunnel.